The rise and evolving role of Fusobacterium nucleatum subspecies
- PMID: 40547902
- PMCID: PMC12178932
- DOI: 10.1016/j.crmicr.2025.100414
The rise and evolving role of Fusobacterium nucleatum subspecies
Abstract
This review examines the classification, distribution, and detection methods of Fusobacterium nucleatum subspecies, focusing on their distinct roles in health and disease. The evolution of F. nucleatum classification is traced from early to modern protein-based and genomic methods, such as 16S rRNA and next generation sequencing and matrix-assisted laser desorption/ionization-time of flight mass spectrometry (MALDI-TOF MS). The review highlights the importance of distinguishing F. nucleatum subspecies due to their varying pathogenic potentials and ecological niches in both oral and extraoral environments. Different subspecies exhibit distinct prevalence and activity levels in specific clinical conditions, such as periodontitis and oral squamous cell carcinoma (OSCC). This highlights the importance of accurately identifying subspecies to understand their role in disease progression. Moreover, understanding the varying pathogenic potential of F. nucleatum subspecies, which is driven by genetic diversity and virulence factors, is also essential for advancing research and improving patient outcomes.
Keywords: Colorectal cancer (CRC); Detection methods; F. nucleatum classification; F. nucleatum subspecies; Oral squamous cell carcinoma (OSCC).
© 2025 The Authors.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures







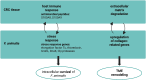
References
-
- Abed J., Emgård J.E.M., Zamir G., Faroja M., Almogy G., Grenov A., Sol A., Naor R., Pikarsky E., Atlan K.A., Mellul A., Chaushu S., Manson A.L., Earl A.M., Ou N., Brennan C.A., Garrett W.S., Bachrach G. Fap2 Mediates Fusobacterium nucleatum colorectal adenocarcinoma enrichment by binding to tumor-expressed gal-GalNAc. Cell Host Microbe. 2016;20:215–225. doi: 10.1016/j.chom.2016.07.006. - DOI - PMC - PubMed
-
- Al-hebshi N.N., Nasher A.T., Maryoud M.Y., Homeida H.E., Chen T., Idris A.M., Johnson N.W. Inflammatory bacteriome featuring Fusobacterium nucleatum and Pseudomonas aeruginosa identified in association with oral squamous cell carcinoma. Sci. Rep. 2017;7:1834. doi: 10.1038/s41598-017-02079-3. - DOI - PMC - PubMed
-
- Allali I., Delgado S., Marron P.I., Astudillo A., Yeh J.J., Ghazal H., Amzazi S., Keku T., Azcarate-Peril M.A. Gut microbiome compositional and functional differences between tumor and non-tumor adjacent tissues from cohorts from the US and Spain. Gut Microbes. 2015;6:161–172. doi: 10.1080/19490976.2015.1039223. - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources

