miR-137 targets Myc to regulate growth during eye development
- PMID: 40554764
- PMCID: PMC12338917
- DOI: 10.1242/dev.204373
miR-137 targets Myc to regulate growth during eye development
Abstract
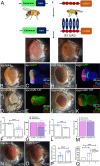
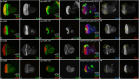
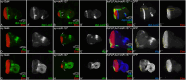
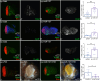
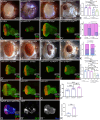
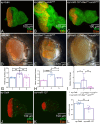
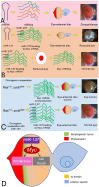
During development, regulation of gene expression is key to cellular homeostasis. Gene expression regulation by non-coding RNAs involves the prevention of mRNA accumulation or the inhibition of translation of their target gene. In a forward-genetic screen to identify the microRNA involved in the growth and patterning of the Drosophila eye, we identified the highly conserved miR-137. Gain of function of miR-137 results in a reduced-eye phenotype by downregulating retinal determination and differentiation markers, and by upregulating negative regulators of eye development, such as Wingless (Wg) and Homothorax (Hth). Loss of function of miR-137 results in an enlarged-eye phenotype. Using bioinformatics and genetic approaches, we identified the oncogene Myc as the target of miR-137. Gain of function of Myc can rescue the reduced-eye phenotype of miR-137 gain of function, and vice versa. We tested the role of miR-137 in regulating Myc levels in the RasV12;scribRNAi, a tumor model of oncogenic cooperation that results in neoplastic tumors. Gain of function of miR-137 in the RasV12;scribRNAi background significantly reduced tumor phenotype as well as Myc levels in the eye. Our studies highlight miR-137 as a post-transcriptional regulator of Myc and a promising therapeutic target for diseases associated with Myc accumulation.
Keywords: Drosophila; Myc; Cell death; Cell proliferation; Eye development; Retina; Retinal determination; miR-137; miRNA.
© 2025. Published by The Company of Biologists.
Conflict of interest statement
Competing interests The authors declare no competing or financial interests.
Figures


Similar articles
-
Regulation of Notch signaling by miR-79 in Drosophila eye development.Biochem Biophys Res Commun. 2025 Aug 30;776:152222. doi: 10.1016/j.bbrc.2025.152222. Epub 2025 Jun 21. Biochem Biophys Res Commun. 2025. PMID: 40561754
-
Targeting miR-32-5p suppresses c-MYC-driven proliferation and induces apoptosis in MCF-7 breast cancer cells.Med Oncol. 2025 Jul 25;42(9):377. doi: 10.1007/s12032-025-02935-7. Med Oncol. 2025. PMID: 40711670 Free PMC article.
-
IGF2 Is Up-regulated by Epigenetic Mechanisms in Hepatocellular Carcinomas and Is an Actionable Oncogene Product in Experimental Models.Gastroenterology. 2016 Dec;151(6):1192-1205. doi: 10.1053/j.gastro.2016.09.001. Epub 2016 Sep 7. Gastroenterology. 2016. PMID: 27614046
-
What a tangled web we weave: crosstalk between JAK-STAT and other signalling pathways during development in Drosophila.FEBS J. 2025 Jul;292(13):3298-3320. doi: 10.1111/febs.17391. Epub 2025 Jan 16. FEBS J. 2025. PMID: 39821459 Review.
-
MicroRNAs as biomarkers in spontaneous intracerebral hemorrhage: A systematic review of recent clinical evidence.Clin Neurol Neurosurg. 2022 Feb;213:107130. doi: 10.1016/j.clineuro.2022.107130. Epub 2022 Jan 14. Clin Neurol Neurosurg. 2022. PMID: 35066247
References
-
- Atkins, M., Potier, D., Romanelli, L., Jacobs, J., Mach, J., Hamaratoglu, F., Aerts, S. and Halder, G. (2016). An ectopic network of transcription factors regulated by Hippo signaling drives growth and invasion of a malignant tumor model. Curr. Biol. 26, 2101-2113. 10.1016/j.cub.2016.06.035 - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

