Overexpression of a Malus baccata (L.) Borkh WRKY Factor Gene MbWRKY33 Increased High Salinity Stress Tolerance in Arabidopsis thaliana
- PMID: 40565296
- PMCID: PMC12193393
- DOI: 10.3390/ijms26125833
Overexpression of a Malus baccata (L.) Borkh WRKY Factor Gene MbWRKY33 Increased High Salinity Stress Tolerance in Arabidopsis thaliana
Abstract
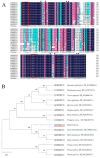
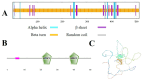
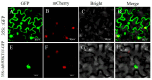
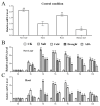
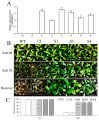
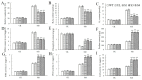
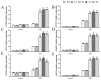
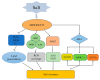
The WRKY transcription factor family is a significant family of plant transcription factors (TFs). Plant growth and development are often influenced by abiotic factors, such as salinity and low temperature. Numerous studies have demonstrated that WRKY TFs primarily influence plant responses to adversity. However, there are few studies on the role of WRKY genes in the stress responses of Malus baccata (L.) Borkh. We cloned the MbWRKY33 gene from Malus baccata for this research, and its roles in salt stress tolerance were analyzed. Phylogenetic tree analysis revealed that MbWRKY33 and PbWRKY33 have the highest homology. Subcellular localization revealed that MbWRKY33 was located within the nucleus. An analysis of tissue-specific expression showed that MbWRKY33 had relatively high expression levels in young leaves and roots. Moreover, Arabidopsis thaliana plants overexpressing MbWRKY33 exhibited stronger resistance to salt stress compared with the wild type (WT) and the unloaded line empty vector (UL). Under the treatment of 200 mM NaCl, transgenic Arabidopsis thaliana plants exhibited significantly higher activities of antioxidant enzymes like superoxide dismutase (SOD), peroxidase (POD), and catalase (CAT) than the control. In contrast, the WT and the UL lines had elevated levels of malondialdehyde (MDA) and reactive oxygen species (ROS). In addition, MbWRKY33 elevates transgenic plant resistance to salt stress by regulating the expression levels of AtNHX1, AtSOS1, AtSOS3, AtNCED3, AtSnRK2, and AtRD29a. Results indicated that MbWRKY33 in Malus might be linked to high-salinity stress responses, laying a foundation for understanding WRKY TFs' reaction to such stress.
Keywords: Malus baccata; MbWRKY33; genetic transformation; high-salinity stress; transcriptional regulation.
Conflict of interest statement
The authors declare no competing interests.
Figures
References
-
- Alscher R.G., Donahue J.L., Cramer C.L. Reactive oxygen species and antioxidants: Relationships in green cells. Physiol. Plant. 1997;100:224–233. doi: 10.1111/j.1399-3054.1997.tb04778.x. - DOI
-
- Han D., Shi Y., Yu Z., Liu W., Lv B., Wang B., Yang G. Isolation and functional analysis of MdCS1: A gene encoding a citrate synthase in Malus domestica (L.) Borkh. Plant Growth Regul. 2015;75:209–218. doi: 10.1007/s10725-014-9945-5. - DOI
-
- Thakur P., Nayyar H. Plant Acclimation to Environmental Stress. Springer; New York, NY, USA: 2013. Facing the Cold Stress by Plants in the Changing Environment: Sensing, Signaling, and Defending Mechanisms; pp. 29–69.
-
- Farooq M., Hussain M., Wakeel A., Siddique K.H.M. Salt stress in maize: Effects, resistance mechanisms, and management. A review. Agron. Sustain. Dev. 2015;35:461–481. doi: 10.1007/s13593-015-0287-0. - DOI
MeSH terms
Substances
Grants and funding
- 32172521/National Natural Science Foundation of China
- 32402458/National Natural Science Foundation of China
- LH2022C097/Natural Science Fund Joint Guidance Project of Heilongjiang Province
- 2023MD744175/China Postdoctoral Science Foundation
- 2022YFD1600501-13/National Key Research and Development Program of China
LinkOut - more resources
Full Text Sources
Miscellaneous

