Adapting to UV: Integrative Genomic and Structural Analysis in Bacteria from Chilean Extreme Environments
- PMID: 40565314
- PMCID: PMC12192789
- DOI: 10.3390/ijms26125842
Adapting to UV: Integrative Genomic and Structural Analysis in Bacteria from Chilean Extreme Environments
Abstract
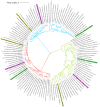
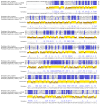
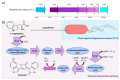
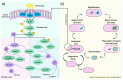
Extremophilic bacteria from extreme environments, such as the Atacama Desert, Salar de Huasco, and Antarctica, exhibit adaptations to intense UV radiation. In this study, we investigated the genomic and structural mechanisms underlying UV resistance in three bacterial isolates identified as Bacillus velezensis PQ169, Pseudoalteromonas sp. AMH3-8, and Rugamonas violacea T1-13. Through integrative genomic analyses, we identified key genes involved in DNA-repair systems, pigment production, and spore formation. Phylogenetic analyses of aminoacidic sequences of the nucleotide excision repair (NER) system revealed conserved evolutionary patterns, indicating their essential role across diverse bacterial taxa. Structural modeling of photolyases from Pseudoalteromonas sp. AMH3-8 and R. violacea T1-13 provided further insights into protein function and interactions critical for DNA repair and UV resistance. Additionally, the presence of a complete violacein operon in R. violacea T1-13 underscores pigment biosynthesis as a crucial protective mechanism. In B. velezensis PQ169, we identified the complete set of genes responsible for sporulation, suggesting that sporulation may represent a key protective strategy employed by this bacterium in response to environmental stress. Our comprehensive approach underscores the complexity and diversity of microbial adaptations to UV stress, offering potential biotechnological applications and advancing our understanding of microbial resilience in extreme conditions.
Keywords: DNA repair; UV resistance; extremophilic bacteria; photolyase; pigment biosynthesis; sporulation.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures


References
-
- Martin-Cuadrado A.B., Senel E., Martínez-García M., Cifuentes A., Santos F., Almansa C., Moreno-Paz M., Blanco Y., García-Villadangos M., García del Cura M.A., et al. Prokaryotic and viral community of the sulfate-rich crust from Peñahueca ephemeral lake, an astrobiology analogue. Environ. Microbiol. 2019;21:3577–3600. doi: 10.1111/1462-2920.14680. - DOI - PubMed
-
- Azua-Bustos A., González-Silva C., Fairén A.G. The Atacama Desert in northern Chile as an analog model of Mars. Front. Astron. Space Sci. 2022;8:810426. doi: 10.3389/fspas.2021.810426. - DOI
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

