Comparative Genomic Analysis Across Multiple Species to Identify Candidate Genes Associated with Important Traits in Chickens
- PMID: 40565519
- PMCID: PMC12192435
- DOI: 10.3390/genes16060627
Comparative Genomic Analysis Across Multiple Species to Identify Candidate Genes Associated with Important Traits in Chickens
Abstract
Background: As one of the most important poultry species worldwide, chickens provide substantial amounts of meat, eggs, and other products for human consumption. With continuous improvements in living standards, consumer demand for high-quality animal products is increasing, making it essential to understand the genetic basis of key traits such as egg production, meat quality, and disease resistance for targeted genetic improvement. Methods: In this study, a number of the candidate genes associated with important traits in chickens were screened by various comparative genomics analysis methods. To further clarify the relationship between these candidate genes and important traits in chickens, they were functionally annotated through the KOG, GO, and KEGG databases. Results: These candidate genes are mainly concentrated in the functional categories of transcription and signal transduction mechanisms and are involved in biological processes such as cyclic nucleotide biosynthesis and intracellular signaling, which involve signaling pathways such as ECM-receptor interactions and calcium signaling. Conclusions: Based on the annotation results from various databases, a functional search of the candidate genes and related literature reports, the following results were obtained: genes such as TBX22, LCORL, and GH were associated with chicken growth traits; genes such as A-FABP, H-FABP, and PRKAB2 were associated with chicken meat quality; genes such as IGF-1, SLC25A29, and WDR25 were associated with chicken reproductive traits; and genes such as C1QBP, VAV2 and IL12B were associated with chicken disease resistance traits. Overall, the findings of this study provide novel insights and candidate genes for genetic improvements in chickens, laying a foundation for future research and breeding strategies targeting key economic traits.
Keywords: candidate genes; chicken; comparative genomics; functional annotation; gene families.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
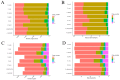
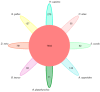

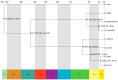
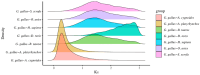

Similar articles
-
Identification of new candidate genes affecting drip loss in pigs based on genomics and transcriptomics data.J Anim Sci. 2025 Jan 4;103:skaf177. doi: 10.1093/jas/skaf177. J Anim Sci. 2025. PMID: 40485044 Free PMC article.
-
Signs and symptoms to determine if a patient presenting in primary care or hospital outpatient settings has COVID-19.Cochrane Database Syst Rev. 2022 May 20;5(5):CD013665. doi: 10.1002/14651858.CD013665.pub3. Cochrane Database Syst Rev. 2022. PMID: 35593186 Free PMC article.
-
A meta-analysis of genome-wide association studies to identify candidate genes associated with feed efficiency traits in pigs.J Anim Sci. 2025 Jan 4;103:skaf010. doi: 10.1093/jas/skaf010. J Anim Sci. 2025. PMID: 39847436 Free PMC article.
-
A rapid and systematic review of the clinical effectiveness and cost-effectiveness of topotecan for ovarian cancer.Health Technol Assess. 2001;5(28):1-110. doi: 10.3310/hta5280. Health Technol Assess. 2001. PMID: 11701100
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2021 Apr 19;4(4):CD011535. doi: 10.1002/14651858.CD011535.pub4. Cochrane Database Syst Rev. 2021. Update in: Cochrane Database Syst Rev. 2022 May 23;5:CD011535. doi: 10.1002/14651858.CD011535.pub5. PMID: 33871055 Free PMC article. Updated.
Cited by
-
Exploring the genetic basis of Newcastle disease virus in chickens: a comprehensive review.Front Immunol. 2025 Jun 27;16:1614794. doi: 10.3389/fimmu.2025.1614794. eCollection 2025. Front Immunol. 2025. PMID: 40655137 Free PMC article. Review.
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

