Beyond Transposons: TIGD1 as a Pan-Cancer Biomarker and Immune Modulator
- PMID: 40565566
- PMCID: PMC12192532
- DOI: 10.3390/genes16060674
Beyond Transposons: TIGD1 as a Pan-Cancer Biomarker and Immune Modulator
Abstract
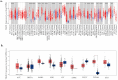
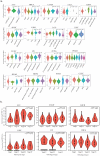
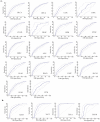
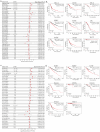
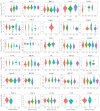
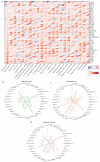
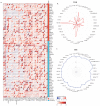
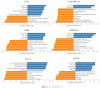
Background/Objectives: TIGD1 (Trigger Transposable Element Derived 1) is a recently identified oncogene with largely unexplored biological functions. Emerging evidence suggests its involvement in multiple cellular processes across cancer types. This study aimed to perform a comprehensive pan-cancer analysis of TIGD1 to evaluate its expression patterns, diagnostic utility, prognostic value, and association with immunotherapy response and drug resistance. Methods: Transcriptomic and clinical data from TCGA and GTEx were analyzed using various bioinformatic tools. Expression profiling, survival analysis, immune correlation studies, gene set enrichment, single-cell sequencing, and drug sensitivity assessments were performed. Results: TIGD1 was found to be significantly upregulated in various tumor types, with notably high expression in colon adenocarcinoma. Elevated TIGD1 expression was associated with poor prognosis in several cancers. TIGD1 levels correlated with key features of the tumor immune microenvironment, including immune checkpoint gene expression, TMB, and MSI, suggesting a role in modulating anti-tumor immunity. GSEA and single-cell analyses implicated TIGD1 in oncogenic signaling pathways. Furthermore, high TIGD1 expression was linked to resistance to several therapeutic agents, including Zoledronate, Dasatinib, and BLU-667. Conclusions: TIGD1 may serve as a promising diagnostic and prognostic biomarker, particularly in colon, gastric, liver, and lung cancers. Its strong associations with immune modulation and therapy resistance highlight its potential as a novel target for precision oncology and immunotherapeutic intervention.
Keywords: TIGD1; cancer prognosis; drug resistance; immune microenvironment; immunotherapy; microsatellite instability; pan-cancer analysis; tumor mutational burden.
Conflict of interest statement
The authors declare no relevant financial or non-financial conflicts of interest.
Figures


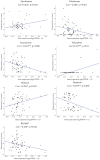
Similar articles
-
Integrated pan-cancer analysis of ADM's role in prognosis, immune modulation and resistance.Front Immunol. 2025 Jun 3;16:1573250. doi: 10.3389/fimmu.2025.1573250. eCollection 2025. Front Immunol. 2025. PMID: 40529377 Free PMC article.
-
Comprehensive pan-cancer analysis reveals NTN1 as an immune infiltrate risk factor and its potential prognostic value in SKCM.Sci Rep. 2025 Jan 25;15(1):3223. doi: 10.1038/s41598-025-85444-x. Sci Rep. 2025. PMID: 39863609 Free PMC article.
-
The Potential of ESCO2 as a Prognostic and Immunotherapeutic Marker of Pan-Cancer and Its Role in Anti-PD-1 Treatment of Bladder Cancer.Oncology. 2025;103(7):610-627. doi: 10.1159/000542188. Epub 2024 Nov 11. Oncology. 2025. PMID: 39527939
-
A systematic review and meta-analysis of long noncoding RNA 00963 expression and prognosis and clinicopathological characteristic in human cancers.Pathol Res Pract. 2023 Feb;242:154291. doi: 10.1016/j.prp.2022.154291. Epub 2022 Dec 26. Pathol Res Pract. 2023. PMID: 36696806
-
Cost-effectiveness of using prognostic information to select women with breast cancer for adjuvant systemic therapy.Health Technol Assess. 2006 Sep;10(34):iii-iv, ix-xi, 1-204. doi: 10.3310/hta10340. Health Technol Assess. 2006. PMID: 16959170
References
-
- Rodić N., Sharma R., Sharma R., Zampella J., Dai L., Taylor M.S., Hruban R.H., Iacobuzio-Donahue C.A., Maitra A., Torbenson M.S., et al. Long Interspersed Element-1 Protein Expression Is a Hallmark of Many Human Cancers. Am. J. Pathol. 2014;184:1280–1286. doi: 10.1016/j.ajpath.2014.01.007. - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

