Genetic Variation and Metapopulation Structure Inform Recovery Goals in a Threatened Species
- PMID: 40565586
- PMCID: PMC12192496
- DOI: 10.3390/genes16060694
Genetic Variation and Metapopulation Structure Inform Recovery Goals in a Threatened Species
Abstract
Background: Monitoring genetic parameters is important for setting effective conservation and management strategies, particularly for small, fragmented, and isolated populations. Small, isolated populations face increased rates of genetic drift and inbreeding, which increase extinction risk especially when gene flow is limited.
Methods: Here, we applied a Genotyping-in-Thousands by sequencing (GT-seq) panel to inform recovery action for the federally threatened northern Idaho ground squirrel (Urocitellus brunneus). We evaluated genetic diversity, structure, connectivity, and effective population size to address species recovery goals.
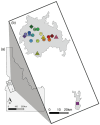
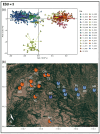
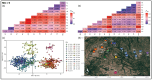
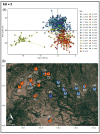
Results: We delineated three types of conservation units: (1) three evolutionarily significant units that represent long-term population structure and variation, (2) nine management units that reflect current demographic connectivity and restrictions to gene flow, and (3) three adaptive units that capture adaptive differentiation across the species range. Effective population sizes per management unit were small overall (mean 38.16, range 2.3-220.9), indicating that recovery goals of 10 subpopulations with Ne > 500 have not been reached.
Conclusions: Our results support the maintenance of connectivity within evolutionarily significant units through the restoration of dispersal corridors. Next steps could include further sampling of some subpopulations with low sample sizes, unsampled subpopulations, and subpopulations that are geographically isolated. Genotyping future samples with the same GT-seq panel would help to detect dispersal, assess effective population size, monitor the effects of inbreeding, and evaluate adaptive differentiation to monitor the effects of management action and environmental change.
Keywords: GT-seq; Urocitellus; conservation units; effective population size; ground squirrels.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

Similar articles
-
Noninvasive sampling reveals landscape genetic structure in the threatened ghost bat (Macroderma gigas) in an ore-rich region of Western Australia.J Hered. 2025 Jul 21;116(4):453-465. doi: 10.1093/jhered/esaf011. J Hered. 2025. PMID: 40067397 Free PMC article.
-
Behavioral interventions to reduce risk for sexual transmission of HIV among men who have sex with men.Cochrane Database Syst Rev. 2008 Jul 16;(3):CD001230. doi: 10.1002/14651858.CD001230.pub2. Cochrane Database Syst Rev. 2008. PMID: 18646068
-
Range-Wide Genomic Analysis Reveals Regional and Meta-Population Dynamics of Decline and Recovery in the Grey Seal.Mol Ecol. 2025 Jul;34(14):e17824. doi: 10.1111/mec.17824. Epub 2025 Jun 19. Mol Ecol. 2025. PMID: 40536212 Free PMC article.
-
Antibody tests for identification of current and past infection with SARS-CoV-2.Cochrane Database Syst Rev. 2022 Nov 17;11(11):CD013652. doi: 10.1002/14651858.CD013652.pub2. Cochrane Database Syst Rev. 2022. PMID: 36394900 Free PMC article.
-
Strategies to improve smoking cessation rates in primary care.Cochrane Database Syst Rev. 2021 Sep 6;9(9):CD011556. doi: 10.1002/14651858.CD011556.pub2. Cochrane Database Syst Rev. 2021. PMID: 34693994 Free PMC article.
References
-
- Allendorf F.W., Funk W.C., Aitken S.N., Byrne M., Luikart G. Conservation and the Genomics of Populations. Oxford University Press; Oxford, UK: 2022.
-
- Allendorf F.W., Luikart G.H., Aitken S.N. Conservation and the Genetics of Populations. Wiley-Blackwell; Malden, MA, USA: 2012.
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

