One-Year Monitoring of the Evolution of SARS-CoV-2 Omicron Subvariants Through Wastewater Analysis (Central Italy, August 2023-July 2024)
- PMID: 40566504
- PMCID: PMC12193976
- DOI: 10.3390/life15060850
One-Year Monitoring of the Evolution of SARS-CoV-2 Omicron Subvariants Through Wastewater Analysis (Central Italy, August 2023-July 2024)
Abstract
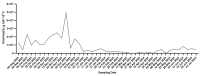
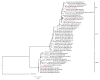
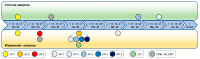
Wastewater surveillance has proven to be a cost-effective, non-invasive method for monitoring the spread and evolution of SARS-CoV-2, yet its value during today's low-incidence phase is still being defined. Between August 2023 and July 2024, 42 composite wastewater samples were collected in Perugia, Italy and analyzed using RT-qPCR and whole-genome sequencing to identify circulating SARS-CoV-2 lineages. In parallel, clinical samples (respiratory tract samples) were collected and analyzed, allowing for direct comparisons to confirm the robustness of the wastewater findings. The sewage viral loads ranged from 8.9 × 105 to 4.9 × 107 genome copies inhabitant-1 day-1, outlining two modest community waves (September-December 2023 and May-July 2024). Sequencing resolved 403 Omicron lineages and revealed three successive subvariant phases: (i) XBB.* dominance (August-October 2023), when late-Omicron XBB subvariants (mainly EG.5.* and XBB.1.5) accounted for almost all genomes; (ii) a BA.2.86/JN surge (November 2023-March 2024), during which the BA.2.86 subvariant, driven mainly by its JN descendants (especially JN.1), rapidly displaced XBB.* and peaked at 89% in February 2024; and (iii) KP.* takeover (April-July 2024), with JN.1-derived KP subvariants rising steadily and KP.3 reaching 81% by July 2024, thereby becoming the dominant lineage. Comparisons of data from wastewater and clinical surveillance demonstrated how the former presented a much higher diversity of circulating viral lineages. Importantly, some subvariants (including BA.2.86*) were detected in wastewater weeks to months prior to clinical identification, and for longer periods. Taken together, the obtained data validated wastewater surveillance as an effective early warning system, especially during periods of low infection prevalence and/or limited molecular testing efforts. This methodology can thus complement clinical surveillance by offering valuable insights into viral dynamics at the community level and enhancing pandemic preparedness.
Keywords: RT-qPCR; SARS-CoV-2; next-generation sequencing; viral variants; wastewater surveillance.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures



Similar articles
-
Monitoring SARS-CoV-2 variants in wastewater during periods of low clinical case surveillance in Ethiopia.mSphere. 2025 Aug 26;10(8):e0022925. doi: 10.1128/msphere.00229-25. Epub 2025 Jul 29. mSphere. 2025. PMID: 40728295 Free PMC article.
-
Alpha to JN.1 variants: SARS-CoV-2 genomic analysis unfolding its various lineages/sublineages evolved in Chhattisgarh, India from 2020 to 2024.World J Virol. 2025 Jun 25;14(2):100001. doi: 10.5501/wjv.v14.i2.100001. World J Virol. 2025. PMID: 40575649 Free PMC article.
-
Long-Term Genomic Surveillance of SARS-CoV‑2 in Campus Wastewater Depicts Lineage Trends and Public Health Implications during and after Omicron Waves.Environ Health (Wash). 2025 Jun 2;3(8):908-919. doi: 10.1021/envhealth.5c00048. eCollection 2025 Aug 15. Environ Health (Wash). 2025. PMID: 40837692 Free PMC article.
-
Rapid, point-of-care antigen tests for diagnosis of SARS-CoV-2 infection.Cochrane Database Syst Rev. 2022 Jul 22;7(7):CD013705. doi: 10.1002/14651858.CD013705.pub3. Cochrane Database Syst Rev. 2022. PMID: 35866452 Free PMC article.
-
A rapid and systematic review of the clinical effectiveness and cost-effectiveness of paclitaxel, docetaxel, gemcitabine and vinorelbine in non-small-cell lung cancer.Health Technol Assess. 2001;5(32):1-195. doi: 10.3310/hta5320. Health Technol Assess. 2001. PMID: 12065068
References
-
- Manirambona E., Okesanya O.J., Olaleke N.O., Oso T.A., Lucero-Prisno D.E. Evolution and Implications of SARS-CoV-2 Variants in the Post-Pandemic Era. Discov. Public Health. 2024;21:16. doi: 10.1186/s12982-024-00140-x. - DOI
-
- Aggiornamento Dei Dati Del Bollettino Della Sorveglianza Integrata COVID-19 in Italia. [(accessed on 24 April 2025)]. Available online: https://www.epicentro.iss.it/coronavirus/aggiornamenti.
-
- Shanmugam B.K., Alqaydi M., Abdisalam D., Shukla M., Santos H., Samour R., Petalidis L., Oliver C.M., Brudecki G., Bin Salem S., et al. A Narrative Review of High Throughput Wastewater Sample Processing for Infectious Disease Surveillance: Challenges, Progress, and Future Opportunities. Int. J. Env. Res. Public Health. 2024;21:1432. doi: 10.3390/ijerph21111432. - DOI - PMC - PubMed
-
- Zhao L., Xu J., Guo J., Zhang P., Guo X., Zuo Z., Gao L., Jia Z., Xue P., Wang J. An Epidemiologic Surveillance Study Based on Wastewater and Respiratory Specimens Reveals Influenza a Virus Prevalence and Mutations in Taiyuan, China during 2023–2024. BMC Infect. Dis. 2024;24:1286. doi: 10.1186/s12879-024-10169-7. - DOI - PMC - PubMed
-
- Annan J., Henderson R., Gray M., Clark R.G., Sarin C., Black K. A Review of Wastewater-Based Epidemiology for the SARS-CoV-2 Virus in Rural, Remote, and Resource-Constrained Settings Internationally: Insights for Implementation, Research, and Policy for First Nations in Canada. Int. J. Env. Res. Public Health. 2024;21:1429. doi: 10.3390/ijerph21111429. - DOI - PMC - PubMed
Grants and funding
- Ricerca Corrente-Linea 1 on emerging and re-emerging infections/Ministero della Salute
- SARS-CoV-2 epidemiological surveillance in urban wastewater (Regional Council resolution n. 1296, 16 December 2021)/Regione Umbria
- Wastewater Virome Analysis for Environmental Surveillance at hospital building level. Next Generation EU, Missione 4 - Componente 2 - LI 1.4 - CUP F53C24001620001/Unione Europea
LinkOut - more resources
Full Text Sources
Miscellaneous

