Evolution of diverse (and advanced) cognitive abilities through adaptive fine-tuning of learning and chunking mechanisms
- PMID: 40566910
- PMCID: PMC12198896
- DOI: 10.1098/rstb.2024.0117
Evolution of diverse (and advanced) cognitive abilities through adaptive fine-tuning of learning and chunking mechanisms
Abstract
The evolution of cognition is frequently discussed as the evolution of cognitive abilities or the evolution of some neuronal structures in the brain. However, since such traits or abilities are often highly complex, understanding their evolution requires explaining how they could have gradually evolved through selection acting on heritable variations in simpler cognitive mechanisms. With this in mind, making use of a previously proposed theory, here, we show how the evolution of cognitive abilities can be captured by the fine-tuning of basic learning mechanisms and, in particular, chunking mechanisms. We use the term chunking broadly for all types of non-elemental learning, claiming that the process by which elements are combined into chunks and associated with other chunks, or elements, is critical for what the brain can do, and that it must be fine-tuned to ecological conditions. We discuss the relevance of this approach to studies in animal cognition, using examples from animal foraging and decision-making, problem-solving and cognitive flexibility. Finally, we explain how even the apparent human-animal gap in sequence learning ability can be explained in terms of different fine-tunings of a similar chunking process.This article is part of the Theo Murphy meeting issue 'Selection shapes diverse animal minds'.
Keywords: animal cognition; cognitive evolution; configural learning; evolution of learning; non-elemental learning; sequence learning.
Conflict of interest statement
We declare we have no competing interests.
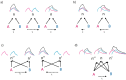
Figures

Similar articles
-
Short-Term Memory Impairment.2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 31424720 Free Books & Documents.
-
Sexual Harassment and Prevention Training.2024 Mar 29. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2024 Mar 29. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 36508513 Free Books & Documents.
-
"In a State of Flow": A Qualitative Examination of Autistic Adults' Phenomenological Experiences of Task Immersion.Autism Adulthood. 2024 Sep 16;6(3):362-373. doi: 10.1089/aut.2023.0032. eCollection 2024 Sep. Autism Adulthood. 2024. PMID: 39371355
-
The Lived Experience of Autistic Adults in Employment: A Systematic Search and Synthesis.Autism Adulthood. 2024 Dec 2;6(4):495-509. doi: 10.1089/aut.2022.0114. eCollection 2024 Dec. Autism Adulthood. 2024. PMID: 40018061 Review.
-
The Black Book of Psychotropic Dosing and Monitoring.Psychopharmacol Bull. 2024 Jul 8;54(3):8-59. Psychopharmacol Bull. 2024. PMID: 38993656 Free PMC article. Review.
Cited by
-
Selection shapes diverse animal minds.Philos Trans R Soc Lond B Biol Sci. 2025 Jun 26;380(1929):20240108. doi: 10.1098/rstb.2024.0108. Epub 2025 Jun 26. Philos Trans R Soc Lond B Biol Sci. 2025. PMID: 40566907 Free PMC article.
-
Sequences and animal intelligence.Philos Trans R Soc Lond B Biol Sci. 2025 Jun 26;380(1929):20240116. doi: 10.1098/rstb.2024.0116. Epub 2025 Jun 26. Philos Trans R Soc Lond B Biol Sci. 2025. PMID: 40566917 Free PMC article. Review.
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

