SELECT: high-precision genome editing strategy via integration of CRISPR-Cas and DNA damage response for cross-species applications
- PMID: 40568938
- PMCID: PMC12199148
- DOI: 10.1093/nar/gkaf595
SELECT: high-precision genome editing strategy via integration of CRISPR-Cas and DNA damage response for cross-species applications
Abstract
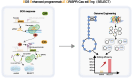
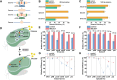
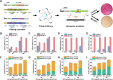
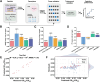
CRISPR-based methods enable genome modifications for diverse applications but often face challenges, such as inconsistent efficiencies, reduced performance in iterative modifications, and difficulties generating high-quality datasets for high-throughput genome engineering. Here, we present SELECT (SOS Enhanced programmabLE CRISPR-Cas ediTing), a novel strategy integrating the CRISPR-Cas system with the DNA damage response. By employing designed and optimized double-strand break induced promoters that are activated upon genome editing, SELECT enables a counter-selection process to eliminate unedited cells, ensuring high-fidelity editing. This approach achieves up to 100% efficiency for point mutations, iterative knockouts, and insertions. In high-throughput library editing, SELECT achieved up to 94.2% efficiency and preserved higher library diversity compared with conventional methods. Application of SELECT in flaviolin biosynthesis resulted in a 3.97-fold increase in production. Furthermore, integration with machine learning tools allowed rapid mapping of genotype-phenotype relationships. SELECT provides a versatile platform for precision genome engineering in Escherichia coli and Saccharomyces cerevisiae.
© The Author(s) 2025. Published by Oxford University Press on behalf of Nucleic Acids Research.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures


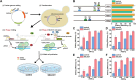
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

