Cell type-specific immune regulation under symbiosis in a facultatively symbiotic coral
- PMID: 40569035
- PMCID: PMC12278269
- DOI: 10.1093/ismejo/wraf132
Cell type-specific immune regulation under symbiosis in a facultatively symbiotic coral
Abstract
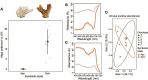
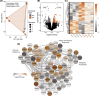
Many cnidarians host single-celled algae within gastrodermal cells, yielding a mutually beneficial exchange of nutrients between host and symbiont, and dysbiosis can lead to host mortality. Previous research has uncovered symbiosis tradeoffs, including suppression of immune pathways in hosts, and correlations between symbiotic state and pathogen susceptibility. Here, we used a multiomic approach to characterize symbiotic states of the facultatively symbiotic coral Oculina arbuscula by generating genotype-controlled fragments of symbiotic and aposymbiotic tissue. 16S rRNA gene sequencing showed no difference in bacterial communities between symbiotic states. Whole-organism proteomics revealed differential abundance of proteins related to immunity, confirming immune suppression during symbiosis. Single-cell RNAseq identified diverse cell clusters within seven cell types across symbiotic states. Specifically, the gastrodermal cell clusters containing algal-hosting cells from symbiotic tissue had higher expression of nitrogen cycling and lipid metabolism genes than aposymbiotic gastrodermal cells. Furthermore, differential enrichment of immune system gene pathways and lower expression of genes involved in immune regulation were observed in these gastrodermal cells from symbiotic tissue. However, there were no differences in gene expression in the immune cell cluster between symbiotic states. We conclude that there is evidence for compartmentalization of immune system regulation in specific gastrodermal cells in symbiosis. This compartmentalization may limit symbiosis tradeoffs by dampening immunity in algal-hosting cells while simultaneously maintaining general organismal immunity.
Keywords: coral; immunity; proteomics; single-cell RNA-sequencing; symbiosis.
© The Author(s) 2025. Published by Oxford University Press on behalf of the International Society for Microbial Ecology.
Conflict of interest statement
None declared.
Figures




References
-
- Lesser MP, Stat M, Gates RD. The endosymbiotic dinoflagellates (Symbiodinium sp.) of corals are parasites and mutualists. Coral Reefs 2013;32:603–11. 10.1007/s00338-013-1051-z - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

