Host-Guest Interactions of Cucurbit[7]uril with Nabumetone and Naproxen: Spectroscopic, Calorimetric, and DFT Studies in Aqueous Solution
- PMID: 40572523
- PMCID: PMC12195925
- DOI: 10.3390/molecules30122558
Host-Guest Interactions of Cucurbit[7]uril with Nabumetone and Naproxen: Spectroscopic, Calorimetric, and DFT Studies in Aqueous Solution
Abstract
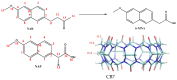
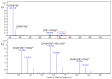
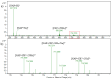
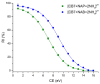
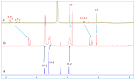
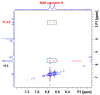
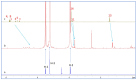
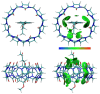
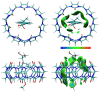
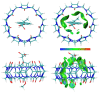
The complexation of nabumetone (NAB) and naproxen (NAP) with cucurbit[7]uril (CB7) was investigated in aqueous solution by isothermal titration microcalorimetry, mass spectrometry, NMR spectroscopy, and computation methods. High-resolution mass spectrometry was used for the determination of the binding stoichiometry and the gas-phase stability of the drug-CB7 complex. The doubly charged NH4+ or Na+ adducts of the 1:1 complex were observed in the mass spectra. The dissociation of complexes was monitored at different collision energies, (1-16) eV, leading to the neutral loss of NH3 and the drug, with charge retention observed on CB7. By performing ITC experiments, all the thermodynamic parameters were determined for the NAB-CB7 complex in water at 25 °C. The corresponding values amounted to the following: logK = 4.66 ± 0.01; ΔrG° = -26.7 ± 0.1 kJ/mol; ΔrH° = -20.2 ± 0.7 kJ/mol; TΔrS° = 6.4 ± 0.8 kJ/mol, i.e., the formation of the inclusion complex is enthalpy driven and has a favorable entropy. The inclusion phenomena were further confirmed by NMR spectroscopy (1H, ROESY, and DOSY), suggesting the encapsulation of the naphthalene ring of both drugs inside the CB7 cavity. The results of the DFT calculations and the IGMH analysis were in accordance with the experimental ones, suggesting that van der Waals interactions play a major role in drug-CB7 complexation.
Keywords: DFT; HR-MS; IGMH; ITC; NMR; cucurbit[7]uril; inclusion complex; nabumetone; naproxen.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Inhibited Hydrolysis of 4-(N,N-Dimethylamino)aryl Imines in a Weak Acid by Supramolecular Encapsulation inside Cucurbit[7]uril.J Org Chem. 2025 Jun 20;90(24):8309-8318. doi: 10.1021/acs.joc.5c00774. Epub 2025 Jun 10. J Org Chem. 2025. PMID: 40493859
-
Supramolecular complexation of locked-flavylium fluorescent dye by cucurbit[7]uril: photophysics and pKa modulation.Phys Chem Chem Phys. 2025 Aug 28;27(34):17877-17883. doi: 10.1039/d5cp02318a. Phys Chem Chem Phys. 2025. PMID: 40785570
-
WITHDRAWN: Non-aspirin, non-steroidal anti-inflammatory drugs for treating osteoarthritis of the knee.Cochrane Database Syst Rev. 2007 Jul 18;2006(1):CD000142. doi: 10.1002/14651858.CD000142.pub2. Cochrane Database Syst Rev. 2007. PMID: 17636601 Free PMC article.
-
Drugs for preventing postoperative nausea and vomiting in adults after general anaesthesia: a network meta-analysis.Cochrane Database Syst Rev. 2020 Oct 19;10(10):CD012859. doi: 10.1002/14651858.CD012859.pub2. Cochrane Database Syst Rev. 2020. PMID: 33075160 Free PMC article.
-
Rofecoxib for rheumatoid arthritis.Cochrane Database Syst Rev. 2005 Jan 25;2005(1):CD003685. doi: 10.1002/14651858.CD003685.pub2. Cochrane Database Syst Rev. 2005. PMID: 15674912 Free PMC article.
References
-
- Steed J.W., Atwood J.L. Supramolecular Chemistry. John Wiley& Sons; Hoboken, NJ, USA: 2022.
-
- Yin H., Wang Z., Wang R. Modulation of Chemical and Biological Properties of Biomedically Relevant Guest Molecules by Cucurbituril-Type Hosts. In: Liu Y., Chen Y., Zhang H.-Y., editors. Handbook of Macrocyclic Supramolecular Assembly. Springer; Singapore: 2019. pp. 1–25. - DOI
LinkOut - more resources
Full Text Sources
Research Materials

