Identification of the Glyceraldehyde-3-Phosphate Dehydrogenase (GeGAPDH) Gene Family in Gastrodia elata Revealing Its Response Characteristics to Low-Temperature and Pathogen Stress
- PMID: 40573855
- PMCID: PMC12197075
- DOI: 10.3390/plants14121866
Identification of the Glyceraldehyde-3-Phosphate Dehydrogenase (GeGAPDH) Gene Family in Gastrodia elata Revealing Its Response Characteristics to Low-Temperature and Pathogen Stress
Abstract
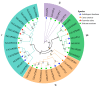
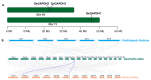
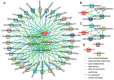
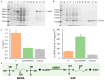
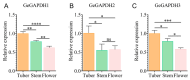
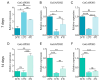
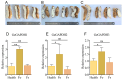
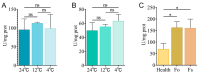
The glyceraldehyde-3-phosphate dehydrogenase (GAPDH) gene plays a pivotal role in the glycolysis/gluconeogenesis process, contributing significantly to glycosyl donor synthesis, plant growth and development, and stress responses. Gastrodia elata Bl., a heterotrophic plant in the Orchidaceae family, has its dried tubers used as the traditional Chinese medicine. This study identified three GeGAPDH genes in G. elata, all encoding basic, stable, hydrophilic proteins. Phylogenetic analysis and subcellular localization predictions categorized GeGAPDH1 as a plastid subtype, while GeGAPDH2 and GeGAPDH3 were classified as cytoplasmic subtypes. Prokaryotic expression experiments demonstrated successful expression of the GeGAPDH1 protein in Escherichia coli, which exhibited significant GAPDH enzymatic activity. Subcellular localization experiments showed that GeGAPDH1 was localized in the plastid. Expression analysis indicated that the three GeGAPDH genes were predominantly expressed in tubers. Under low-temperature stress, although the total GAPDH enzyme activity in tubers did not change significantly, the expression of GeGAPDH1 was significantly up-regulated, while GeGAPDH2 and GeGAPDH3 were significantly down-regulated. This suggests that different subtypes of GeGAPDH may regulate cold resistance through different pathways. Upon pathogen infection, the GeGAPDH gene family exhibited pathogen-specific regulatory patterns. During infection by Fusarium oxysporum, both the expression levels of all three GeGAPDH genes and the total GAPDH enzyme activity in tubers increased significantly; however, F. solani infection induced a significant increase in total GAPDH enzyme activity without significant changes in gene expression. These results suggest that the GeGAPDH gene family may respond to different pathogen infections through transcriptional or translational regulation mechanisms. This study systematically identified and characterized the GeGAPDH gene family in G. elata, providing a theoretical foundation for understanding the functional differentiation of GAPDH in heterotrophic plants.
Keywords: Gastrodia elata; enzymatic activity; glyceraldehyde-3-phosphate dehydrogenase; stress response; subcellular localization.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures





Similar articles
-
Draft Genome and Biological Characteristics of Fusarium solani and Fusarium oxysporum Causing Black Rot in Gastrodia elata.Int J Mol Sci. 2023 Feb 25;24(5):4545. doi: 10.3390/ijms24054545. Int J Mol Sci. 2023. PMID: 36901977 Free PMC article.
-
Genome-Wide Identification and Characterisation of the 4-Coumarate-CoA Ligase (4CL) Gene Family in Gastrodia elata and Their Transcriptional Response to Fungal Infection.Int J Mol Sci. 2025 Aug 6;26(15):7610. doi: 10.3390/ijms26157610. Int J Mol Sci. 2025. PMID: 40806741 Free PMC article.
-
First Report of Leaf Spot Caused by Curvularia buchloes on Polygonatum sibiricum in China.Plant Dis. 2025 Jun 26. doi: 10.1094/PDIS-12-24-2725-PDN. Online ahead of print. Plant Dis. 2025. PMID: 40570357
-
Antiretrovirals for reducing the risk of mother-to-child transmission of HIV infection.Cochrane Database Syst Rev. 2011 Jul 6;(7):CD003510. doi: 10.1002/14651858.CD003510.pub3. Cochrane Database Syst Rev. 2011. PMID: 21735394
-
Levetiracetam add-on for drug-resistant focal epilepsy: an updated Cochrane Review.Cochrane Database Syst Rev. 2012 Sep 12;2012(9):CD001901. doi: 10.1002/14651858.CD001901.pub2. Cochrane Database Syst Rev. 2012. PMID: 22972056 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

