Frequencies and subtypes of glycophorin GYP(B-A-B) hybrids among northern Thais, Burmese, and Karen with a previous history of malaria infection: a study in the Thailand-Myanmar border area
- PMID: 40574735
- PMCID: PMC12199742
- DOI: 10.7717/peerj.19589
Frequencies and subtypes of glycophorin GYP(B-A-B) hybrids among northern Thais, Burmese, and Karen with a previous history of malaria infection: a study in the Thailand-Myanmar border area
Abstract
Background: Evidence indicates that genetic variations in the GYP(B-A-B) hybrid genes are associated with protection against malaria. Therefore, this study aims to characterize the GYP(B-A-B) hybrid alleles among northern Thais, Burmese, and Karen with and without a previous history of malaria infection.
Methods: A total of 709 DNA samples were genotyped to identify GYP(B-A-B) hybrids using PCR-sequence specific primers (PCR-SSP) combined with Sanger sequencing. Additionally, some DNA samples (n = 243) were also tested with high-resolution melting (HRM) analysis.
Results: In our sampled populations, 14/87 (16.0%), 3/34 (8.8%), 0/16 (0%), and 1/18 (5.6%) of northern Thais, Burmese, Karen, and other minorities in Myanmar with a previous history of malaria infection, respectively, were identified with GYP(B-A-B) hybrid genes, whereas individuals without a history of malaria infection were 24/155 (15.5%), 5/183 (2.6%), and 4/216 (1.9%) in northern Thais, Burmese, and Karen, respectively. In the latter groups, DNA sequences showed that 17/155 (11.0%) northern Thais were GYP*Mur/GYPB heterozygotes and the other 6/155 (3.9%) were GYP*Thai/GYPB heterozygotes. The remaining one (0.6%) sample was a GYP*Mur/GYP*Mur homozygote. Among Burmese, 3/183 (1.6%) were GYP*Mur/GYPB heterozygotes and 1/183 (0.5%) was GYP*Thai/GYPB heterozygote. The remaining one (0.5%) sample being a GYP*Mur/GYP*Mur homozygote. Among Karen samples, all four were GYP*Mur/GYPB heterozygotes.
Conclusion: Across all studied populations, GYP*Mur was the predominant allele, followed by GYP*Thai. In addition, genotyping results obtained by HRM were consistent with PCR-SSP combined with Sanger sequencing. A statistically non-significant association was noted for the glycophorin GYP(B-A-B) hybrids and malaria infection. Our findings provide insight into genetic variations of GYP(B-A-B) hybrid alleles among populations in the Thailand-Myanmar border area. This information could be used as a guideline to identify compatible blood products for transfusion and to prevent alloimmunization.
Keywords: Blood group genotyping; GYP(B-A-B) hybrid alleles; Malaria; Mia antigen.
©2025 Srichankhot et al.
Conflict of interest statement
The authors declare there are no competing interests.
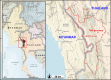
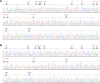
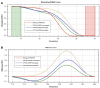
Figures


Similar articles
-
Characterization of GYP(B-A-B) hybrid glycophorins among Thai blood donors with Mia-positive phenotypes.Blood Transfus. 2024 May;22(3):198-205. doi: 10.2450/BloodTransfus.567. Epub 2023 Oct 24. Blood Transfus. 2024. PMID: 38063789 Free PMC article.
-
Molecular genetic analysis of Mia -positive hybrid glycophorins revealed two novel alleles of GP.Vw and multiple variant transcripts of GYPB existing in both the homozygous GP.Mur and wild-type GPB individuals.Transfusion. 2021 Aug;61(8):2477-2486. doi: 10.1111/trf.16450. Epub 2021 Jun 12. Transfusion. 2021. PMID: 34117642
-
Molecular and hematological characteristics of two different δ-globin promoter variants, δ-276(A>G) and δ-77(T >C) among Thai, Burmese, and Laotian subjects.PeerJ. 2025 Jul 3;13:e19636. doi: 10.7717/peerj.19636. eCollection 2025. PeerJ. 2025. PMID: 40620769 Free PMC article.
-
Indoor residual spraying for preventing malaria in communities using insecticide-treated nets.Cochrane Database Syst Rev. 2022 Jan 17;1(1):CD012688. doi: 10.1002/14651858.CD012688.pub3. Cochrane Database Syst Rev. 2022. PMID: 35038163 Free PMC article.
-
Signs and symptoms to determine if a patient presenting in primary care or hospital outpatient settings has COVID-19.Cochrane Database Syst Rev. 2022 May 20;5(5):CD013665. doi: 10.1002/14651858.CD013665.pub3. Cochrane Database Syst Rev. 2022. PMID: 35593186 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

