The genome sequence of the greater argentine, Argentina silus (Ascanius, 1775)
- PMID: 40574744
- PMCID: PMC12198749
- DOI: 10.12688/wellcomeopenres.23691.1
The genome sequence of the greater argentine, Argentina silus (Ascanius, 1775)
Abstract
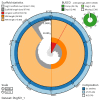
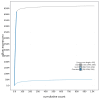
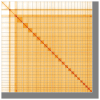
We present a genome assembly from a male specimen of Argentina silus (the greater argentine; Chordata; Actinopteri; Argentiniformes; Argentinidae). The genome sequence has a total length of 670.70 megabases. Most of the assembly is scaffolded into 24 chromosomal pseudomolecules, including the X and Y sex chromosomes. The mitochondrial genome has also been assembled and is 16.64 kilobases in length. Gene annotation of this assembly on Ensembl identified 19,422 protein-coding genes.
Keywords: Argentina silus; Argentiniformes; chromosomal; genome sequence; greater argentine.
Copyright: © 2025 Vang HBM et al.
Conflict of interest statement
No competing interests were disclosed.
Figures


References
-
- Árnason E: A report on the mitochondrial DNA variation of the great silver smelt. Argentina silus,2017.
LinkOut - more resources
Full Text Sources

