The emergence of NY10: insights into the 2012 West Nile Virus outbreak in the United States
- PMID: 40574749
- PMCID: PMC12202104
- DOI: 10.1093/ve/veaf037
The emergence of NY10: insights into the 2012 West Nile Virus outbreak in the United States
Abstract
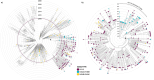
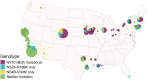
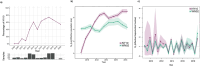
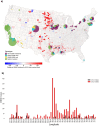
West Nile Virus (WNV) remains a public health risk across North America due to its capacity for rapid adaptation and evolution. While research in the United States has focused on the WN02 and SW03 mutations, the NY10 genotype, first detected in 2010, has received comparatively little attention. We conducted a phylogenetic and phylodynamic investigation of NY10, revealing its rapid increase in detection frequency and effective population size in the early 2010s. Our analysis suggests that NY10 played an important role in the 2012 WNV outbreak, with an effective population size indicating higher diversity than other lineages during this period. Despite this, NY10 appears geographically restricted, with no detections west of Colorado, indicating that barriers in the southwestern United States may influence its spread. These findings highlight the complex interplay between viral evolution, geography, and the environmental factors that shape WNV epidemiology. The study emphasizes the potential of WNV to generate genotypes with epidemic potential and underscores the importance of integrating genetic data into surveillance and forecasting systems to better predict and manage future outbreaks. Understanding the drivers of WNV's genetic diversity will be crucial for developing more effective public health strategies as the virus continues to evolve.
Keywords: NY10; WNV; West Nile virus; arbovirus; phylogenetic analysis.
Published by Oxford University Press 2025.
Figures

Similar articles
-
Accreditation through the eyes of nurse managers: an infinite staircase or a phenomenon that evaporates like water.J Health Organ Manag. 2025 Jun 30. doi: 10.1108/JHOM-01-2025-0029. Online ahead of print. J Health Organ Manag. 2025. PMID: 40574247
-
Early warning systems and rapid response systems for the prevention of patient deterioration on acute adult hospital wards.Cochrane Database Syst Rev. 2021 Nov 22;11(11):CD005529. doi: 10.1002/14651858.CD005529.pub3. Cochrane Database Syst Rev. 2021. PMID: 34808700 Free PMC article.
-
Carbamazepine versus phenytoin monotherapy for epilepsy: an individual participant data review.Cochrane Database Syst Rev. 2017 Feb 27;2(2):CD001911. doi: 10.1002/14651858.CD001911.pub3. Cochrane Database Syst Rev. 2017. Update in: Cochrane Database Syst Rev. 2019 Jul 18;7:CD001911. doi: 10.1002/14651858.CD001911.pub4. PMID: 28240353 Free PMC article. Updated.
-
Drugs for preventing postoperative nausea and vomiting in adults after general anaesthesia: a network meta-analysis.Cochrane Database Syst Rev. 2020 Oct 19;10(10):CD012859. doi: 10.1002/14651858.CD012859.pub2. Cochrane Database Syst Rev. 2020. PMID: 33075160 Free PMC article.
-
Signs and symptoms to determine if a patient presenting in primary care or hospital outpatient settings has COVID-19.Cochrane Database Syst Rev. 2022 May 20;5(5):CD013665. doi: 10.1002/14651858.CD013665.pub3. Cochrane Database Syst Rev. 2022. PMID: 35593186 Free PMC article.
References
-
- Andersen-lab . west-nile-genomics-wnv4k. andersen-lab/west-nile-genomics-wnv4k, 2023. 2019-03-28T21:12:15Z.
LinkOut - more resources
Full Text Sources

