Genetic and codon usage analyses reveal the evolution of the seoul virus
- PMID: 40574799
- PMCID: PMC12198216
- DOI: 10.3389/fgene.2025.1544577
Genetic and codon usage analyses reveal the evolution of the seoul virus
Abstract
Introduction: Seoul virus (Orthohantavirus seoulense, SEOV), a member of the Hantaviridae, causes hemorrhagic fever with renal syndrome (HFRS) through rodent hosts. However, its molecular evolutionary dynamics and codon usage patterns remain poorly understood.
Methods: This study integrated coding sequences from GenBank and previously acquired SEOV strains to systematically analyze genetic evolution and codon usage bias.
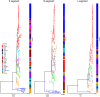
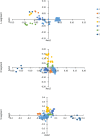
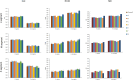
Results: It revealed that SEOV evolved seven clades (A-G) with distinct amino acid variation sites and geographic clustering. Recombination events were identified during evolution, alongside purifying and positive selection on specific sites (e.g., codon 259 in the S segment and codon 11 in the M segment). The three viral segments (L, M, and S) exhibited weak codon usage bias, predominantly driven by natural selection, with host adaptation significantly influencing evolutionary trajectories. The S segment demonstrated the strongest pathogenicity due to its closer codon usage alignment with Homo sapiens (H. sapiens) and Rattus norvegicus (R. norvegicus), whereas the L segment showed the lowest host adaptation. Divergent codon preferences among clades highlighted adaptive strategies in host-virus interactions.
Conclusion: These findings elucidate the evolutionary mechanisms of SEOV and provide a theoretical foundation for live attenuated vaccine design and region-specific viral control strategies.
Keywords: codon usage bias; evolution; genetic; hantavirus; phylogenetic; seoul virus.
Copyright © 2025 Wei, Cai, Han, Han, Zhang, Xu, Jiang and Li.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Natural Selection as the Primary Driver of Codon Usage Bias in the Mitochondrial Genomes of Three Medicago Species.Genes (Basel). 2025 May 30;16(6):673. doi: 10.3390/genes16060673. Genes (Basel). 2025. PMID: 40565565 Free PMC article.
-
Comparative analysis of codon usage bias and host adaptation across avian metapneumovirus genotypes.Poult Sci. 2025 Jun 11;104(9):105428. doi: 10.1016/j.psj.2025.105428. Online ahead of print. Poult Sci. 2025. PMID: 40541102 Free PMC article.
-
Sequential versus standard triple first-line therapy for Helicobacter pylori eradication.Cochrane Database Syst Rev. 2016 Jun 28;2016(6):CD009034. doi: 10.1002/14651858.CD009034.pub2. Cochrane Database Syst Rev. 2016. PMID: 27351542 Free PMC article.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2021 Apr 19;4(4):CD011535. doi: 10.1002/14651858.CD011535.pub4. Cochrane Database Syst Rev. 2021. Update in: Cochrane Database Syst Rev. 2022 May 23;5:CD011535. doi: 10.1002/14651858.CD011535.pub5. PMID: 33871055 Free PMC article. Updated.
-
Molecular evolution and reassortment dynamics of Orthohantavirus hantanense revealed through longitudinal genomic surveillance in the Republic of Korea.Sci Rep. 2025 Jul 9;15(1):24672. doi: 10.1038/s41598-025-11003-z. Sci Rep. 2025. PMID: 40634677 Free PMC article.
References
LinkOut - more resources
Full Text Sources

