Novel perspectives on plastome evolution in Onagraceae
- PMID: 40574894
- PMCID: PMC12190799
- DOI: 10.1093/aobpla/plaf025
Novel perspectives on plastome evolution in Onagraceae
Abstract
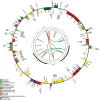
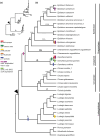
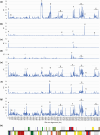
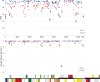
Previous systematic studies have generated abundant information on plants in family Onagraceae Juss., making this taxonomic group a model for understanding plant evolution. The chloroplast genome is widely used to provide valuable insights into how plant lineages evolved. In the present study, we employed shotgun sequencing to assemble new plastomes from Onagraceae. Plastomes of ten species and one genus, Fuchsia, are reported for the first time. We characterize and compare the plastome features of six genera (Chamaenerion, Circaea, Epilobium, Fuchsia, Ludwigia, and Oenothera), allowing us to reconstruct their phylogenies and explore inter- and infra-generic evolutionary relationships, inverted repeat (IR) expansion, plastome size increases, and correlations among repeat elements, genetic variations, and evolutionary events. Our findings indicate that each of the tribes and subfamilies we assessed exhibits unique plastome features. Our phylogenetic tree supports previous findings, but also reveals that some clades need further systematic analyses. We show that increased plastome size within subfamily Onagroideae coincides with IR expansion, which is not the case for subfamily Ludwigioideae. In addition, our results indicate that higher repeat numbers and greater genetic variation can serve as indicators of evolutionary events, such as gene loss and gain, IR boundary shifts, and inversions, but they may not have arisen universally across all members of Onagraceae. Our study provides some novel insights into plastome evolution in the Onagraceae. Further studies should aim to elucidate how plastome size has evolved in Ludwigioideae and explore the evolutionary roles of regions in Onagraceae plastomes exhibiting high repeat numbers and genetic variations.
© The Author(s) 2025. Published by Oxford University Press on behalf of the Annals of Botany Company.
Conflict of interest statement
None declared.
Figures

Similar articles
-
The 2 Sigma Genus Concept in mammalogy: Lessons from Lasiurus.PLoS One. 2025 Jun 25;20(6):e0325554. doi: 10.1371/journal.pone.0325554. eCollection 2025. PLoS One. 2025. PMID: 40560834 Free PMC article.
-
Signs and symptoms to determine if a patient presenting in primary care or hospital outpatient settings has COVID-19.Cochrane Database Syst Rev. 2022 May 20;5(5):CD013665. doi: 10.1002/14651858.CD013665.pub3. Cochrane Database Syst Rev. 2022. PMID: 35593186 Free PMC article.
-
Factors that influence parents' and informal caregivers' views and practices regarding routine childhood vaccination: a qualitative evidence synthesis.Cochrane Database Syst Rev. 2021 Oct 27;10(10):CD013265. doi: 10.1002/14651858.CD013265.pub2. Cochrane Database Syst Rev. 2021. PMID: 34706066 Free PMC article.
-
A rapid and systematic review of the clinical effectiveness and cost-effectiveness of paclitaxel, docetaxel, gemcitabine and vinorelbine in non-small-cell lung cancer.Health Technol Assess. 2001;5(32):1-195. doi: 10.3310/hta5320. Health Technol Assess. 2001. PMID: 12065068
-
Comparative Analysis of Plastomes of Artemisia and Insights into the Infra-Generic Phylogenetic Relationships Within the Genus.Genes (Basel). 2025 May 29;16(6):659. doi: 10.3390/genes16060659. Genes (Basel). 2025. PMID: 40565551 Free PMC article.
References
-
- Andrews S. FastQC: a quality control tool for high throughput sequence data. 2010. http://www.bioinformatics.babraham.ac.uk/projects/fastqc
-
- Barloy-Hubler F, Le Gac AL, Boury C et al. Sequencing, de novo assembly of Ludwigia plastomes, and comparative analysis within the Onagraceae family. Peer Community Journal. Section: Genomics 2025;5:e43. https://doi.org/ 10.24072/pcjournal.536 - DOI
LinkOut - more resources
Full Text Sources

