Comprehensive immune ageing reveals TREM2/TIM3 myeloid cells drive brain immune evasion
- PMID: 40580577
- PMCID: PMC12266526
- DOI: 10.1016/j.ebiom.2025.105833
Comprehensive immune ageing reveals TREM2/TIM3 myeloid cells drive brain immune evasion
Abstract
Background: Ageing-dependent low-grade inflammation is a hallmark of central nervous system (CNS) diseases. Vascular and immune abnormalities are implicated in the progression of gliomas and occur in the early stages of Alzheimer's disease (AD); however, the mechanisms by which these alterations manifest in the brain parenchyma remain unclear.
Methods: Using RNAseq, scRNAseq, bioinformatics tools and a cohort of patients with glioma and Alzheimer's disease for validation of results, we have established an analysis of blood-brain barrier (BBB) dysfunction and neuron loss. A mouse model for glioblastoma pathology was also used that reversed BBB disruption and neuron loss, with the incorporation of the IDH mutation. Finally, we established a characterization of the relevant immune populations with an IHC analysis and transcriptional profile.
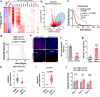
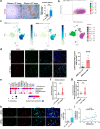
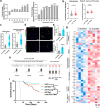
Findings: In this study, molecular analyses of the brain ecosystem revealed that blood-brain barrier dysfunction and neuronal synapse integrity exhibit significant threshold-dependent changes that correlate directly and inversely, respectively, with brain ageing (significant changes at 57 years) and the progression of AD and gliomas (survival of 1525 vs 4084 days for patients with High vs Low BBB dysfunction). Using human samples and mouse models, we identified immunoageing processes characterized by an imbalance between pro-inflammatory and anti-inflammatory signals. This dysregulation promotes the extravasation of monocyte-derived macrophages (85% increase of cells), particularly those with a suppressive phenotype, alongside an increase in inflammatory cytokine levels. Notably, our data show that vascular normalization in a glioma model can reverse neuronal loss and attenuate the aggressiveness of the tumours. Finally, tumour development can be prevented by reactivating the ageing immune system.
Interpretation: We propose that the ageing brain represents a common, BBB dysfunction-associated process driving chronic inflammation. This inflammation is regulated by TREM2+/TIM3+ suppressive myeloid cells, which play a central role in disease progression. Our findings suggest that targeting these pathways could offer therapeutic strategies to mitigate CNS pathologies linked to ageing, characterized by toxic neuroinflammation and myeloid dysfunction.
Funding: This study was funded by ISCIII and co-funded by the European Union.
Keywords: Ageing; Alzheimer's disease; Glioblastoma; Immune evasion; TIM3; TREM2.
Copyright © 2025 The Author(s). Published by Elsevier B.V. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no conflict of interest.
Figures





References
-
- Mogilenko D.A., Shpynov O., Andhey P.S., et al. Comprehensive profiling of an ageing immune system reveals clonal GZMK. Immunity. 2021;54(1):99–115.e12. - PubMed
-
- Segura-Collar B., Mata-Martinez P., Hernandez-Lain A., Sanchez-Gomez P., Gargini R. Blood-brain barrier disruption: a common driver of central nervous system diseases. Neuroscientist. 2022;28(3):222–237. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials

