Leveraging machine learning models in evaluating ADMET properties for drug discovery and development
- PMID: 40585410
- PMCID: PMC12205928
- DOI: 10.5599/admet.2772
Leveraging machine learning models in evaluating ADMET properties for drug discovery and development
Abstract
Background and purpose: The evaluation of ADMET properties remains a critical bottleneck in drug discovery and development, contributing significantly to the high attrition rate of drug candidates. Traditional experimental approaches are often time-consuming, cost-intensive, and limited in scalability. This review aims to investigate how recent advances in machine learning (ML) models are revolutionizing ADMET prediction by enhancing accuracy, reducing experimental burden, and accelerating decision-making during early-stage drug development.
Experimental approach: This article systematically examines the current landscape of ML applications in ADMET prediction, including the types of algorithms employed, common molecular descriptors and datasets used, and model development workflows. It also explores public databases, model evaluation metrics, and regulatory considerations relevant to computational toxicology. Emphasis is placed on supervised and deep learning techniques, model validation strategies, and the challenges of data imbalance and model interpretability.
Key results: ML-based models have demonstrated significant promise in predicting key ADMET endpoints, outperforming some traditional quantitative structure - activity relationship (QSAR) models. These approaches provide rapid, cost-effective, and reproducible alternatives that integrate seamlessly with existing drug discovery pipelines. Case studies discussed in this review illustrate the successful deployment of ML models for solubility, permeability, metabolism, and toxicity predictions.
Conclusion: Machine learning has emerged as a transformative tool in ADMET prediction, offering new opportunities for early risk assessment and compound prioritization. While challenges such as data quality, algorithm transparency, and regulatory acceptance persist, continued integration of ML with experimental pharmacology holds the potential to substantially improve drug development efficiency and reduce late-stage failures.
Keywords: ADMET prediction; AI/ML; computational toxicology; molecular descriptors; pharmacokinetics.
Copyright © 2025 by the authors.
Conflict of interest statement
Conflict of interest: The authors declare no conflicts of interest related to this review article.
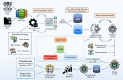
Figures


Similar articles
-
Applications of machine learning in high-entropy alloys: a review of recent advances in design, discovery, and characterization.Nanoscale. 2025 Aug 27. doi: 10.1039/d5nr01562f. Online ahead of print. Nanoscale. 2025. PMID: 40878184 Review.
-
The role of machine learning in predictive toxicology: A review of current trends and future perspectives.Life Sci. 2025 Oct 1;378:123821. doi: 10.1016/j.lfs.2025.123821. Epub 2025 Jun 24. Life Sci. 2025. PMID: 40571275 Review.
-
The dawn of a new era: can machine learning and large language models reshape QSP modeling?J Pharmacokinet Pharmacodyn. 2025 Jun 16;52(4):36. doi: 10.1007/s10928-025-09984-5. J Pharmacokinet Pharmacodyn. 2025. PMID: 40524056 Free PMC article. Review.
-
A systematic review of automated prediction of sudden cardiac death using ECG signals.Physiol Meas. 2025 Jan 23;13(1). doi: 10.1088/1361-6579/ad9ce5. Physiol Meas. 2025. PMID: 39657316 Review.
-
Cost-effectiveness of using prognostic information to select women with breast cancer for adjuvant systemic therapy.Health Technol Assess. 2006 Sep;10(34):iii-iv, ix-xi, 1-204. doi: 10.3310/hta10340. Health Technol Assess. 2006. PMID: 16959170
References
-
- Berdigaliyev N., Aljofan M.. An overview of drug discovery and development. Future Medicinal Chemistry 12 (2020) 939-947. https://doi.org/10.4155/fmc-2019-0307 10.4155/fmc-2019-0307 - DOI - PubMed
-
- Pei Z.. Computer-aided drug discovery: From traditional simulation methods to language models and quantum computing. Cell Reports Physical Science 5 (2024) 11-21. https://doi.org/10.1016/j.xcrp.2024.102334 10.1016/j.xcrp.2024.102334 - DOI
-
- Vamathevan J., Clark D., Czodrowski P., Dunham I., Ferran E., Lee G., Li B., Madabhushi A., Shah P., Spitzer M., Zhao S.. Applications of machine learning in drug discovery and development. Nature Reviews Drug Discovery 18 (2019) 463-477. https://doi.org/10.1038/s41573-019-0024-5 10.1038/s41573-019-0024-5 - DOI - PMC - PubMed
-
- Dara S., Dhamercherla S., Jadav S.S., Babu C.M., Ahsan M.J.. Machine Learning in Drug Discovery: A Review. Artificial Intelligence Review 55 (2022) 1947-1999. https://doi.org/10.1007/s10462-021-10058-4 10.1007/s10462-021-10058-4 - DOI - PMC - PubMed
-
- Xiong G., Wu Z., Yi J., Fu L., Yang Z., Hsieh C., Yin M., Zeng X., Wu C., Lu A., Chen X., Hou T., Cao D.. ADMETlab 2.0: An integrated online platform for accurate and comprehensive predictions of ADMET properties. Nucleic Acids Research 49 (2021) W5-W14. https://doi.org/10.1093/nar/gkab255 10.1093/nar/gkab255 - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources
