FAK signaling suppression by OCT4-ITGA6 mediates the effectively removal of residual pluripotent stem cells and enhances application safety
- PMID: 40585989
- PMCID: PMC12204078
- DOI: 10.7150/thno.111198
FAK signaling suppression by OCT4-ITGA6 mediates the effectively removal of residual pluripotent stem cells and enhances application safety
Abstract
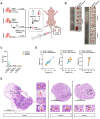
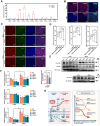
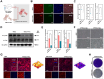
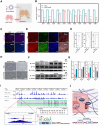
Rationale: Pluripotent stem cells (PSCs) serve as a critical source of seed cells for regenerative therapies due to their unlimited proliferative capacity and ability to differentiate into all three germ layers. Despite their potential, the risk of teratoma formation caused by residual PSCs within differentiated cell populations poses a significant barrier to clinical applications. This study aims to develop a novel strategy to selectively remove residual PSCs while preserving the safety and functionality of PSC-derived differentiated cells (iDCs). Methods: The calcium- and magnesium-free balanced salt solution (BSS(Ca-Mg-)) was employed to selectively target PSCs in a co-culture system comprising PSCs and four types of iDCs. The effect of BSS(Ca-Mg-) treatment on teratoma formation was evaluated in immunodeficient mice following cell transplantation. Comparative analysis and gene knockdown experiments were conducted to explore the molecular mechanisms underlying the differential response of PSCs and iDCs to BSS(Ca-Mg-), focusing on FAK signaling and its interaction with OCT4 and ITGA6. Results: The BSS(Ca-Mg-) treatment effectively induced the detachment of PSCs in the co-culture system without disrupting iDC adhesion. In vivo experiments confirmed that cells treated with BSS(Ca-Mg-) did not form teratomas upon implantation into immunodeficient mice. Mechanistic studies revealed that PSCs exhibit lower activation of FAK signaling compared to iDCs, contributing to their selective detachment. Additionally, OCT4 and ITGA6 were found to maintain each other's protein expression, forming a feedback loop that suppressed FAK signaling, while FAK suppression further enhanced OCT4 expression. Conclusions: The study presents a safe, effective, and cost-efficient method for the selective removal of residual PSCs. This approach enhances existing safety measures for iDC applications, improving the clinical feasibility of iDC-based cell therapies.
Keywords: Pluripotent stem cells; Teratoma; balanced salt solution; focal adhesion kinase; integrin.
© The author(s).
Conflict of interest statement
Competing Interests: Author Jian Wang, Shixin Gong, Zongmin Jiang, Huiyuan Zhang, Lida Wu, and Yuchun Gu are employed by Allife Medicine Inc. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Notch3 enhances the synergistic effect of all-trans retinoic acid and calcipotriol in pancreatic stellate cell activation.J Transl Med. 2025 Jun 22;23(1):694. doi: 10.1186/s12967-025-06666-1. J Transl Med. 2025. PMID: 40545532 Free PMC article.
-
Teratoma Assay for Testing Pluripotency and Malignancy of Stem Cells: Insufficient Reporting and Uptake of Animal-Free Methods-A Systematic Review.Int J Mol Sci. 2023 Feb 15;24(4):3879. doi: 10.3390/ijms24043879. Int J Mol Sci. 2023. PMID: 36835305 Free PMC article.
-
Quality improvement strategies for diabetes care: Effects on outcomes for adults living with diabetes.Cochrane Database Syst Rev. 2023 May 31;5(5):CD014513. doi: 10.1002/14651858.CD014513. Cochrane Database Syst Rev. 2023. PMID: 37254718 Free PMC article.
-
Interventions for eye movement disorders due to acquired brain injury.Cochrane Database Syst Rev. 2018 Mar 5;3(3):CD011290. doi: 10.1002/14651858.CD011290.pub2. Cochrane Database Syst Rev. 2018. PMID: 29505103 Free PMC article.
-
Stem cell transplantation for induction of remission in medically refractory Crohn's disease.Cochrane Database Syst Rev. 2022 May 13;5(5):CD013070. doi: 10.1002/14651858.CD013070.pub2. Cochrane Database Syst Rev. 2022. PMID: 35556242 Free PMC article.
References
-
- Yamanaka S. Pluripotent Stem Cell-Based Cell Therapy-Promise and Challenges. Cell stem cell. 2020;27:523–31. - PubMed
-
- Wang J, Sun S, Deng H. Chemical reprogramming for cell fate manipulation: Methods, applications, and perspectives. Cell stem cell. 2023;30:1130–47. - PubMed
-
- Thomson JA, Itskovitz-Eldor J, Shapiro SS, Waknitz MA, Swiergiel JJ, Marshall VS. et al. Embryonic stem cell lines derived from human blastocysts. Science (New York, NY) 1998;282:1145–7. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

