Quantifying complexity in DNA structures with high resolution Atomic Force Microscopy
- PMID: 40593613
- PMCID: PMC12216981
- DOI: 10.1038/s41467-025-60559-x
Quantifying complexity in DNA structures with high resolution Atomic Force Microscopy
Abstract
DNA topology is essential for regulating cellular processes and maintaining genome stability, yet it is challenging to quantify due to the size and complexity of topologically constrained DNA molecules. By combining high-resolution Atomic Force Microscopy (AFM) with a new high-throughput automated pipeline, we can quantify the length, conformation, and topology of individual complex DNA molecules with sub-molecular resolution. Our pipeline uses deep-learning methods to trace the backbone of individual DNA molecules and identify crossing points, efficiently determining which segment passes over which. We use this pipeline to determine the structure of stalled replication intermediates from Xenopus egg extracts, including theta structures and late replication products, and the topology of plasmids, knots and catenanes from the E. coli Xer recombination system. We use coarse-grained simulations to quantify the effect of surface immobilisation on twist-writhe partitioning. Our pipeline opens avenues for understanding how fundamental biological processes are regulated by DNA topology.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures




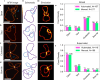

References
-
- Bates, A. D. & Maxwell, A. DNA Topology. (Oxford University Press, Oxford; New York, 2005).
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

