Decoding polyubiquitin regulation of KV7. 1 (KCNQ1) surface expression with engineered linkage-selective deubiquitinases
- PMID: 40593673
- PMCID: PMC12218124
- DOI: 10.1038/s41467-025-60893-0
Decoding polyubiquitin regulation of KV7. 1 (KCNQ1) surface expression with engineered linkage-selective deubiquitinases
Abstract
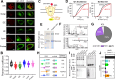
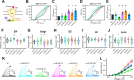
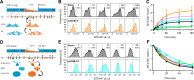
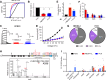
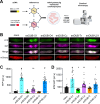
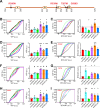
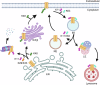
Polyubiquitin chain diversity generates a 'ubiquitin code' that universally regulates protein abundance, localization, and function. Functions of polyubiquitin diversity are mostly unknown, with lack of progress due to an inability to selectively tune protein polyubiquitin linkages in live cells. We develop linkage-selective engineered deubiquitinases (enDUBs) by fusing linkage-selective DUB catalytic domains to GFP-targeted nanobody and use them to investigate polyubiquitin linkage regulation of an ion channel, YFP-KCNQ1. YFP-KCNQ1 in HEK293 cells has polyubiquitin chains with K48/K63 linkages dominant. EnDUBs yield unique effects on channel surface abundance with a pattern indicating: K11 promotes ER retention/degradation, enhances endocytosis, and reduces recycling; K29/K33 promotes ER retention/degradation; K63 enhances endocytosis and reduces recycling; and K48 is necessary for forward trafficking. EnDUB effects differ in cardiomyocytes and on KCNQ1 disease mutants, emphasizing ubiquitin code mutability. The results reveal distinct polyubiquitin chains control different aspects of KCNQ1 abundance and subcellular localization and introduce linkage-selective enDUBs as potent tools to demystify the polyubiquitin code.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: S.A.K. and H.M.C. are inventors on a patent held by Columbia University for “Compositions and methods for using engineered deubiquitinases for probing ubiquitin-dependent cellular processes.” H.M.C. is a scientific co-founder and on the SAB of two startups, Stablix, Inc. and Flux Therapeutics, pursuing targeted protein stabilization therapeutics. S.A.K. is a co-founder and employee of Stablix, Inc. The remaining authors declare no competing interests.
Figures


Update of
-
Decoding polyubiquitin regulation of KV7. 1 functional expression with engineered linkage-selective deubiquitinases.bioRxiv [Preprint]. 2024 Sep 17:2024.09.17.613539. doi: 10.1101/2024.09.17.613539. bioRxiv. 2024. Update in: Nat Commun. 2025 Jul 1;16(1):5805. doi: 10.1038/s41467-025-60893-0. PMID: 39345403 Free PMC article. Updated. Preprint.
References
MeSH terms
Substances
Grants and funding
- S10 RR027050/RR/NCRR NIH HHS/United States
- R01 NS126850/NS/NINDS NIH HHS/United States
- 1F30 HL140878/U.S. Department of Health & Human Services | NIH | National Heart, Lung, and Blood Institute (NHLBI)
- T32 GM007367/GM/NIGMS NIH HHS/United States
- S10RR027050/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
- T32 GM008464/GM/NIGMS NIH HHS/United States
- RO1 HL121253/U.S. Department of Health & Human Services | NIH | National Heart, Lung, and Blood Institute (NHLBI)
- RO1 HL142111/U.S. Department of Health & Human Services | NIH | National Heart, Lung, and Blood Institute (NHLBI)
- P01 HL164319/HL/NHLBI NIH HHS/United States
- R01 HL121253/HL/NHLBI NIH HHS/United States
- POST1019343/American Heart Association (American Heart Association, Inc.)
- F30 HL140878/HL/NHLBI NIH HHS/United States
- R01 HL142111/HL/NHLBI NIH HHS/United States
- P30 CA013696/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources

