Bioinformatic and genomic analysis identifies C allele of APOE rs7412 as the most prominent variant limiting extreme human longevity
- PMID: 40594060
- PMCID: PMC12215757
- DOI: 10.1038/s41598-025-02848-5
Bioinformatic and genomic analysis identifies C allele of APOE rs7412 as the most prominent variant limiting extreme human longevity
Abstract
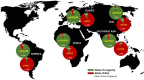
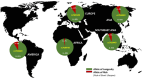
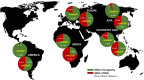
The genetic factors influencing human longevity have long been a subject of curiosity, linking ancient historical records with modern genomic research. This study seeks to identify key genetic contributors to lifespan by integrating genome-wide association studies (GWAS) with functional annotations across multiple biological databases. Among our findings, APOE emerged as one of the most prominent gene, with the rs7412-C variant showing a strong association with reduced lifespan due to its widely distributed across global populations. Additionally, we observed that rs449647-A and rs405509-T were more frequently found in regions with longer life expectancies, such as Asia, and were less common in African populations, suggesting a possible genetic influence on regional lifespan variations. These findings provide a basis for future research into the genetic mechanisms underlying aging and may contribute to the development of strategies to promote healthy aging.
Keywords: APOE; Bioinformatics; GWAS; Longevity; SNP.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no competing interests. Ethics approval and consent to participate: For research involving secondary data analysis of public datasets, consent and ethical clearance were deemed not applicable, as the data were anonymized and publicly available for research use.
Figures




Similar articles
-
Behavioral interventions to reduce risk for sexual transmission of HIV among men who have sex with men.Cochrane Database Syst Rev. 2008 Jul 16;(3):CD001230. doi: 10.1002/14651858.CD001230.pub2. Cochrane Database Syst Rev. 2008. PMID: 18646068
-
A meta-analysis of genome-wide association studies to identify candidate genes associated with feed efficiency traits in pigs.J Anim Sci. 2025 Jan 4;103:skaf010. doi: 10.1093/jas/skaf010. J Anim Sci. 2025. PMID: 39847436 Free PMC article.
-
Measures implemented in the school setting to contain the COVID-19 pandemic.Cochrane Database Syst Rev. 2022 Jan 17;1(1):CD015029. doi: 10.1002/14651858.CD015029. Cochrane Database Syst Rev. 2022. Update in: Cochrane Database Syst Rev. 2024 May 2;5:CD015029. doi: 10.1002/14651858.CD015029.pub2. PMID: 35037252 Free PMC article. Updated.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2021 Apr 19;4(4):CD011535. doi: 10.1002/14651858.CD011535.pub4. Cochrane Database Syst Rev. 2021. Update in: Cochrane Database Syst Rev. 2022 May 23;5:CD011535. doi: 10.1002/14651858.CD011535.pub5. PMID: 33871055 Free PMC article. Updated.
-
Diagnostic test accuracy and cost-effectiveness of tests for codeletion of chromosomal arms 1p and 19q in people with glioma.Cochrane Database Syst Rev. 2022 Mar 2;3(3):CD013387. doi: 10.1002/14651858.CD013387.pub2. Cochrane Database Syst Rev. 2022. PMID: 35233774 Free PMC article.
References
-
- Caroll, R. & Prickett, S. The Bible: Authorized King James Version 1st edn. (Oxford University Press, 2008).
-
- Haleem, M. A. The Qur’an (Oxford World’s Classics) (Oxford University Press, 2008).
-
- Musa, A. Y. A thousand years, less fifty: Toward a quranic view of extreme longevity. In Religion and the Implications of Radical Life Extension (eds Maher, D. F. & Mercer, C.) 123–131 (Palgrave Macmillan US, 2009). 10.1057/9780230100725_11.
-
- Smulders, L. & Deelen, J. Genetics of human longevity: From variants to genes to pathways. J. Intern. Med.295(4), 416–435 (2024). - PubMed
-
- Miyagi, S., Iwama, N., Kawabata, T. & Hasegawa, K. Longevity and diet in Okinawa, Japan: The past, present and future. Asia Pacific J. Public Health15(1_suppl), S3–S9. 10.1177/101053950301500S03 (2003). - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

