Molecular docking and biological evaluation of a novel IWS1 inhibitor for the treatment of human retroperitoneal liposarcoma
- PMID: 40594510
- PMCID: PMC12216911
- DOI: 10.1038/s41598-025-07215-y
Molecular docking and biological evaluation of a novel IWS1 inhibitor for the treatment of human retroperitoneal liposarcoma
Abstract
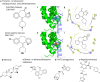
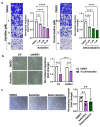
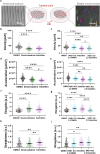
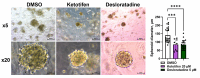
IWS1 is a key assembly factor of the RNA polymerase II (RNAPII) elongation complex, and its overexpression is associated with worse outcomes in patients with liposarcoma (LPS). This study aimed to identify compounds that can disrupt the IWS1/Spt6 interaction and assess their biological effects in dedifferentiated LPS (DDLPS). Using the AlphaFold-predicted structure of IWS1, we identified a core binding region (AA 545-694) for its interaction with Spt6. Through molecular modeling and virtual screening, Ketotifen and Desloratadine were predicted as candidate inhibitors. Both were predicted to mimic Spt6 phenylalanine (F217) and disrupt the complex, which was confirmed by co-immunoprecipitation. Functional assays showed that treatment with either compound reduced migration, invasion, and spheroid formation in DDLPS cell lines. Additionally, increased nuclear localization of IWS1 was observed. These findings suggest Ketotifen and Desloratadine as promising inhibitors of the IWS1/Spt6 interaction, with potential applications in reducing the invasive properties of human LPS.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no competing interests.
Figures


References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

