BMAL1 and ARNT enable circadian HIF2α responses in clear cell renal cell carcinoma
- PMID: 40595592
- PMCID: PMC12215836
- DOI: 10.1038/s41467-025-60904-0
BMAL1 and ARNT enable circadian HIF2α responses in clear cell renal cell carcinoma
Abstract
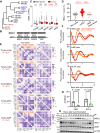
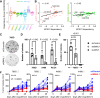
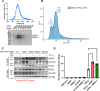
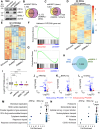
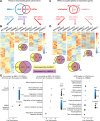
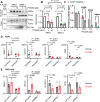
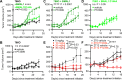
Circadian disruption enhances cancer risk, and many tumors exhibit disordered circadian gene expression. We show rhythmic gene expression is unexpectedly robust in clear cell renal cell carcinoma (ccRCC). The core circadian transcription factor BMAL1 is closely related to ARNT, and we show that BMAL1-HIF2α regulates a subset of HIF2α target genes in ccRCC cells. Depletion of BMAL1 selectively reduces HIF2α chromatin association and target gene expression and reduces ccRCC growth in culture and in xenografts. Analysis of pre-existing data reveals higher BMAL1 in patient-derived xenografts that are sensitive to growth suppression by a HIF2α antagonist (PT2399). BMAL1-HIF2α is more sensitive than ARNT-HIF2α is to suppression by PT2399, and the effectiveness of PT2399 for suppressing xenograft tumor growth in vivo depends on the time of day at which it is delivered. Together, these findings indicate that an alternate HIF2α heterodimer containing the circadian partner BMAL1 influences HIF2α activity, growth, and sensitivity to HIF2α antagonist drugs in ccRCC cells.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures

Update of
-
BMAL1-HIF2α heterodimers contribute to ccRCC.Res Sq [Preprint]. 2024 Jul 16:rs.3.rs-4651047. doi: 10.21203/rs.3.rs-4651047/v1. Res Sq. 2024. Update in: Nat Commun. 2025 Jul 1;16(1):5834. doi: 10.1038/s41467-025-60904-0. PMID: 39070610 Free PMC article. Updated. Preprint.
References
-
- Linehan, W. M. & Ricketts, C. J. The Cancer Genome Atlas of renal cell carcinoma: findings and clinical implications. Nat. Rev. Urol.16, 539–552 (2019). - PubMed
MeSH terms
Substances
Grants and funding
- UL1TR002550/U.S. Department of Health & Human Services | NIH | National Center for Advancing Translational Sciences (NCATS)
- CA271500/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
- R01 CA211187/CA/NCI NIH HHS/United States
- R01 CA271500/CA/NCI NIH HHS/United States
- 450149205/Deutsche Forschungsgemeinschaft (German Research Foundation)
- UL1 TR002550/TR/NCATS NIH HHS/United States
- DK136780/U.S. Department of Health & Human Services | NIH | National Institute of Diabetes and Digestive and Kidney Diseases (National Institute of Diabetes & Digestive & Kidney Diseases)
- CA211187/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
LinkOut - more resources
Full Text Sources
Medical

