The molecular basis for acetylhistidine synthesis by HisAT/NAT16
- PMID: 40595645
- PMCID: PMC12219266
- DOI: 10.1038/s41467-025-61145-x
The molecular basis for acetylhistidine synthesis by HisAT/NAT16
Abstract
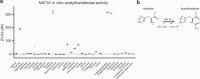
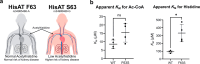
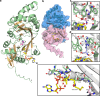
Acetylhistidine has been detected in human blood, but its origin and function are not known. It is formed when the acetyl group of acetyl-CoA is transferred to the α-amino group of histidine. Here we identify the intracellular NAT16 as the human histidine acetyltransferase (HisAT) responsible for histidine acetylation in vitro and in vivo. A NAT16 variant (p.Phe63Ser) present in over 5% of the population was previously found to correlate with reduced plasma levels of acetylhistidine and increased risk of kidney disease. Our biochemical analysis of HisAT/NAT16 Phe63Ser shows reduced affinity for Histidine supporting a model where this variant has less acetylhistidine catalysis leading to lower blood level of acetylhistidine. We find that HisAT adopts a double-GNAT (Gcn5-related N-Acetyltransferase) fold where the N-terminal domain binds acetyl-CoA and with distinct active site conformation allowing the binding of histidine in between the two domains. We detect similar structures from across living organisms and find that the HisAT structure is conserved in several archaeal and bacterial species. In sum, NAT16 is the human histidine acetyltransferase utilizing a rare double-GNAT structure to steer plasma acetylhistidine levels with potential impact for kidney function.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures



Similar articles
-
An ectotherm homologue of human predicted gene NAT16 encodes histidine N-acetyltransferase responsible for Nα-acetylhistidine synthesis.Biochim Biophys Acta. 2014 Jan;1840(1):434-42. doi: 10.1016/j.bbagen.2013.10.004. Epub 2013 Oct 9. Biochim Biophys Acta. 2014. PMID: 24121108
-
Biochemical and structural characterization of a GNAT superfamily protein acetyltransferase from Helicobacter pylori.J Biol Chem. 2025 Jul;301(7):110356. doi: 10.1016/j.jbc.2025.110356. Epub 2025 Jun 10. J Biol Chem. 2025. PMID: 40505863 Free PMC article.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2021 Apr 19;4(4):CD011535. doi: 10.1002/14651858.CD011535.pub4. Cochrane Database Syst Rev. 2021. Update in: Cochrane Database Syst Rev. 2022 May 23;5:CD011535. doi: 10.1002/14651858.CD011535.pub5. PMID: 33871055 Free PMC article. Updated.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2017 Dec 22;12(12):CD011535. doi: 10.1002/14651858.CD011535.pub2. Cochrane Database Syst Rev. 2017. Update in: Cochrane Database Syst Rev. 2020 Jan 9;1:CD011535. doi: 10.1002/14651858.CD011535.pub3. PMID: 29271481 Free PMC article. Updated.
-
Immunogenicity and seroefficacy of pneumococcal conjugate vaccines: a systematic review and network meta-analysis.Health Technol Assess. 2024 Jul;28(34):1-109. doi: 10.3310/YWHA3079. Health Technol Assess. 2024. PMID: 39046101 Free PMC article.
References
-
- Wade, A. M. & Tucker, H. N. Antioxidant characteristics of L-histidine. J. Nutr. Biochem.9, 308–315 (1998).
-
- Thalacker-Mercer, A. E. & Gheller, M. E. Benefits and adverse effects of histidine supplementation. J. Nutr.150, 2588S–2592S (2020). - PubMed
MeSH terms
Substances
Grants and funding
- 772039 (To T. Arnesen)/EC | EU Framework Programme for Research and Innovation H2020 | H2020 Priority Excellent Science | H2020 European Research Council (H2020 Excellent Science - European Research Council)
- 772039/EC | EU Framework Programme for Research and Innovation H2020 | H2020 Priority Excellent Science | H2020 European Research Council (H2020 Excellent Science - European Research Council)
LinkOut - more resources
Full Text Sources

