Repurposing of CNS accumulating drugs Gemfibrozil and Doxylamine for enhanced sensitization of glioblastoma cells through modulation of autophagy
- PMID: 40595852
- PMCID: PMC12219411
- DOI: 10.1038/s41598-025-05054-5
Repurposing of CNS accumulating drugs Gemfibrozil and Doxylamine for enhanced sensitization of glioblastoma cells through modulation of autophagy
Abstract
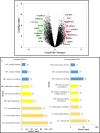
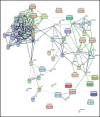
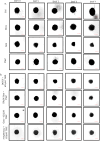
GBM is one of the most aggressive malignancies, having the greatest fatality rate and average life years lost. The current standard medicine, temozolomide (TMZ), is ineffective, requiring the development of new treatments. However, identifying and introducing a novel medicine takes time and money. In this context, repurposing FDA-approved drugs can be a novel yet efficient alternative method. Here, we, therefore, investigated the differential expression signatures of genes of patients suffering from GBM from publicly available GEO datasets and constructed a connectivity map. Functional annotation and KEGG pathway analysis showed dysregulated molecular activities and pathways. Based on their gene ontologies, putative key genes and hub genes linked with the disease were identified, and the C-MAP database was scanned for FDA-approved medicinal compounds that could alter hub gene expression or associated pathways. Our in-silico investigation showed that Gemfibrozil (Gem) and Doxylamine (Doxy) might reverse GBM disease patterns by deregulating GBM-related genes. Evaluation of the GBM inhibitory potential of these drugs through in-vitro and three-dimensional spheroid assay showed promising results. These drugs were more cytotoxic than TMZ; however, they synergised with TMZ as well. Interestingly, the cellular homeostatic process autophagy which has been implicated significantly in GBM pathogenesis and therapy resistance, was found to be inhibited by the drugs Gemfibrozil and Doxylamine, signifying their prospective potential. Therefore, in this study, we, for the first time, identify drugs with the ability to cross the blood brain barrier (BBB), with potential cytotoxic effects beyond TMZ, and with autophagy inhibitory potential, which can be further explored for repurposing against GBM.
Keywords: Autophagy; Disease reversal; Drug repurposing; Glioblastoma multiform; Temozolomide.
© 2025. The Author(s).
Conflict of interest statement
Declararions. Competing interests: The authors declare no competing interests.
Figures








Similar articles
-
Treatment options for progression or recurrence of glioblastoma: a network meta-analysis.Cochrane Database Syst Rev. 2021 May 4;5(1):CD013579. doi: 10.1002/14651858.CD013579.pub2. Cochrane Database Syst Rev. 2021. PMID: 34559423 Free PMC article.
-
A novel approach to enhance glioblastoma multiforme treatment efficacy: non-coding RNA targeted therapy and adjuvant approaches.Clin Epigenetics. 2025 Jun 21;17(1):108. doi: 10.1186/s13148-025-01900-5. Clin Epigenetics. 2025. PMID: 40544306 Free PMC article. Review.
-
Deciphering pericyte-induced temozolomide resistance in glioblastoma with a 3D microphysiological system mimicking the biomechanical properties of brain tissue.Acta Biomater. 2025 Jun 15;200:202-217. doi: 10.1016/j.actbio.2025.05.038. Epub 2025 May 16. Acta Biomater. 2025. PMID: 40383349
-
A novel nuclear RNA HSD52 scaffolding NONO/SFPQ complex modulates DNA damage repair to facilitate temozolomide resistance.Neuro Oncol. 2025 May 15;27(4):963-978. doi: 10.1093/neuonc/noae272. Neuro Oncol. 2025. PMID: 39673809
-
Methadone-mediated sensitization of glioblastoma cells is drug and cell line dependent.J Cancer Res Clin Oncol. 2021 Mar;147(3):779-792. doi: 10.1007/s00432-020-03485-3. Epub 2020 Dec 14. J Cancer Res Clin Oncol. 2021. PMID: 33315125 Free PMC article.
References
-
- Maher, E. A. & Bachoo, R. M. Glioblastoma (2014). 10.1016/B978-0-12-410529-4.00078-4
-
- Burnet, N. G. et al. Clinical and practical considerations for the use of intensity- modulated radiotherapy and image guidance in neuro-oncology statement of search strategies used and general considerations of IMRT for central nervous system tumours. Clin. Oncol.26, 395–406. 10.1016/j.clon.2014.04.024 (2014). - PubMed
-
- Lassman, A. B. Molecular biology of gliomas. Curr. Neurol. Neurosci. Rep.4(3), 228–233. 10.1007/s11910-004-0043-3 (2004). - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials

