MTCH2 Deficiency Promotes E2F4/TFRC-Mediated Ferroptosis and Sensitizes Colorectal Cancer Liver Metastasis to Sorafenib
- PMID: 40600459
- PMCID: PMC12463053
- DOI: 10.1002/advs.202500019
MTCH2 Deficiency Promotes E2F4/TFRC-Mediated Ferroptosis and Sensitizes Colorectal Cancer Liver Metastasis to Sorafenib
Abstract
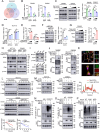
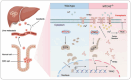
Ferroptosis is a specific type of lipid peroxide-mediated cell death which is crucial in tumor suppression. While the mitochondrial carrier homolog 2 (MTCH2) is implicated in lipid homeostasis and mitochondrial metabolism, its role in ferroptosis and colorectal cancer (CRC) remains uncharacterized. Here, MTCH2 is identified as a crucial regulator of ferroptosis in CRC progression. Clinically, high expression of MTCH2 in CRC tissues predicts poor prognosis. Functionally, loss of MTCH2 inhibits azoxymethane (AOM)/dextran sodium sulfate (DSS)-induced colorectal tumorigenesis in MTCH2cKO mice and leads to accumulation of ferrous ion and enhances ferroptosis of CRC in vitro and in vivo. Mechanistically, MTCH2 deficiency promotes the proteasome-dependent ubiquitination of E2F4 and attenuates transcriptional inhibition of transferrin receptor (TFRC) by E2F4, ultimately facilitating TFRC-mediated ferroptosis in CRC cells. Moreover, MTCH2 depletion combined with sorafenib treatment synergistically triggers ferroptosis, suppresses liver metastasis, and effectively eradicates tumors in liver metastasis foci. Taken together, This study reveals the mechanism of MTCH2 deficiency-induced ferroptosis to inhibit the progression of CRC and supports a potential therapeutic strategy targeting the MTCH2/E2F4/TFRC signaling axis in CRC patients with liver metastasis.
Keywords: MTCH2; colorectal cancer; ferroptosis; liver metastasis; sorafenib.
© 2025 The Author(s). Advanced Science published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures







References
-
- Sung H., Ferlay J., Siegel R. L., Laversanne M., Soerjomataram I., Jemal A., Bray F., CA Cancer J. Clin. 2021, 71, 209. - PubMed
-
- Carlomagno C., De Stefano A., Rosanova M., De Falco S., Attademo L., Fiore G., De Placido S., Cancer Metastasis Rev. 2019, 38, 307. - PubMed
-
- Yu Q., Zhang N., Gan X., Chen L., Wang R., Liang R., Jian J., Phytomedicine 2023, 119, 154999. - PubMed
MeSH terms
Substances
Grants and funding
- 82173218 82171720/National Natural Science Foundation of China
- GZC20230142/Postdoctoral Fellowship Program of CPSF
- 2024M760148/China Postdoctoral Science Foundation
- 2025 Open Project-5/Key Laboratory of Carcinogenesis and Translational Research, Ministry of Education
- ZY202503/Science Foundation of Peking University Cancer Hospital
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous
