Nasopharyngeal microbiome composition by SARS-CoV-2 presence and severity
- PMID: 40603926
- PMCID: PMC12223233
- DOI: 10.1038/s41598-025-01764-y
Nasopharyngeal microbiome composition by SARS-CoV-2 presence and severity
Abstract
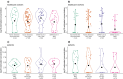
The influence of SARS-CoV-2 on the nasopharyngeal microbiome, or vice-versa, is unclear. Nasopharyngeal swabs from Dutch healthcare workers (N = 257) and hospital outpatients with respiratory symptoms (N = 143), leftover after SARS-CoV-2 testing in 2020-2021, were 16S rRNA amplicon sequenced and tested for respiratory viruses by multiplex PCR panel. The healthcare workers were younger and much healthier than the patients, and experienced less severe viral infections. In the healthcare workers, log10 estimated concentrations (ECs) of Corynebacterium were slightly increased in samples with SARS-CoV-2 versus no virus detected, regardless of symptomatology (adjusted regression coefficient 0.52, p = 0.042) but no other bacterial ECs differed. Corynebacterium and Dolosigranulum ECs were higher in very mild/asymptomatic SARS-CoV-2 episodes compared to very mild/asymptomatic episodes with no viruses detected, but lower in mild compared to very mild/asymptomatic SARS-CoV-2 episodes (-1.07, p = 0.015, and -1.37, p = 0.011, respectively). In the patients, similar but non-significant trends by SARS-CoV-2 severity (fatal, severe, moderate versus mild) were seen for Dolosigranulum, but not for Corynebacterium. In this population, the largest nasopharyngeal microbiome composition differences were seen by the presence and severity of comorbidities. These findings suggest that the Dolosigranulum EC decreases with increasing SARS-CoV-2 severity, but the clinical relevance of this finding is unclear.
Keywords: 16S rRNA gene sequencing; COVID-19; Nasopharyngeal microbiome; Netherlands; SARS-CoV-2.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no competing interests.
Figures



Similar articles
-
Signatures of COVID-19 Severity and Immune Response in the Respiratory Tract Microbiome.mBio. 2021 Aug 31;12(4):e0177721. doi: 10.1128/mBio.01777-21. Epub 2021 Aug 17. mBio. 2021. PMID: 34399607 Free PMC article.
-
Age-Related Changes in the Nasopharyngeal Microbiome Are Associated With Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Infection and Symptoms Among Children, Adolescents, and Young Adults.Clin Infect Dis. 2022 Aug 24;75(1):e928-e937. doi: 10.1093/cid/ciac184. Clin Infect Dis. 2022. PMID: 35247047 Free PMC article.
-
The nasal microbiome modulates risk for SARS-CoV-2 infection.EBioMedicine. 2025 May;115:105660. doi: 10.1016/j.ebiom.2025.105660. Epub 2025 Apr 9. EBioMedicine. 2025. PMID: 40210576 Free PMC article.
-
Nirmatrelvir combined with ritonavir for preventing and treating COVID-19.Cochrane Database Syst Rev. 2023 Nov 30;11(11):CD015395. doi: 10.1002/14651858.CD015395.pub3. Cochrane Database Syst Rev. 2023. PMID: 38032024 Free PMC article.
-
Signs and symptoms to determine if a patient presenting in primary care or hospital outpatient settings has COVID-19.Cochrane Database Syst Rev. 2022 May 20;5(5):CD013665. doi: 10.1002/14651858.CD013665.pub3. Cochrane Database Syst Rev. 2022. PMID: 35593186 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

