Electron flow in hydrogenotrophic methanogens under nickel limitation
- PMID: 40604290
- PMCID: PMC12350162
- DOI: 10.1038/s41586-025-09229-y
Electron flow in hydrogenotrophic methanogens under nickel limitation
Abstract
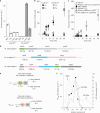
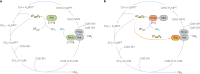
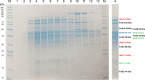
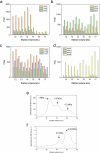
Methanogenic archaea are the main producers of the potent greenhouse gas methane1,2. In the methanogenic pathway from CO2 and H2 studied under laboratory conditions, low-potential electrons for CO2 reduction are generated by a flavin-based electron-bifurcation reaction catalysed by heterodisulfide reductase (Hdr) complexed with the associated [NiFe]-hydrogenase (Mvh)3-5. F420-reducing [NiFe]-hydrogenase (Frh) provides electrons to the methanogenic pathway through the electron carrier F420 (ref. 6). Here we report that under strictly nickel-limited conditions, in which the nickel concentration is similar to those often observed in natural habitats7-11, the production of both [NiFe]-hydrogenases in Methanothermobacter marburgensis is strongly downregulated. The Frh reaction is substituted by a coupled reaction with [Fe]-hydrogenase (Hmd), and the role of Mvh is taken over by F420-dependent electron-donating proteins (Elp). Thus, Hmd provides all electrons for the reducing metabolism under these nickel-limited conditions. Biochemical and structural characterization of Elp-Hdr complexes confirms the electronic interaction between Elp and Hdr. The conservation of the genes encoding Elp and Hmd in CO2-reducing hydrogenotrophic methanogens suggests that the Hmd system is an alternative pathway for electron flow in CO2-reducing hydrogenotrophic methanogens under nickel-limited conditions.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures






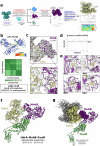

References
-
- Conrad, R. The global methane cycle: recent advances in understanding the microbial processes involved. Environ. Microbiol. Rep.1, 285–292 (2009). - PubMed
-
- Thauer, R. K., Kaster, A. K., Seedorf, H., Buckel, W. & Hedderich, R. Methanogenic archaea: ecologically relevant differences in energy conservation. Nat. Rev. Microbiol.6, 579–591 (2008). - PubMed
-
- Setzke, E., Hedderich, R., Heiden, S. & Thauer, R. K. H2: heterodisulfide oxidoreductase complex from Methanobacterium thermoautotrophicum: composition and properties. Eur. J. Biochem.220, 139–148 (1994). - PubMed
-
- Wagner, T., Koch, J., Ermler, U. & Shima, S. Methanogenic heterodisulfide reductase (HdrABC-MvhAGD) uses two noncubane [4Fe-4S] clusters for reduction. Science357, 699–702 (2017). - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

