In evolution's unending race: ancestral STING sensors in Salmo salar mediate intracellular bacterial detection and programmed cell death through evolutionarily conserved pathways
- PMID: 40607404
- PMCID: PMC12213456
- DOI: 10.3389/fimmu.2025.1570871
In evolution's unending race: ancestral STING sensors in Salmo salar mediate intracellular bacterial detection and programmed cell death through evolutionarily conserved pathways
Abstract
Introduction: "In evolution's unending race, survival demands continuous adaptation- to stop is to fall behind." The Stimulator of Interferon Genes (STING) pathway embodies this principle, acting as a conserved master regulator of cytosolic DNA sensing from Drosophila to salmon and humans. Although extensively characterized in mammals, its structural features and regulatory roles during intracellular bacterial infection in teleosts remain poorly defined.
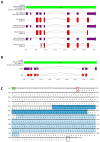
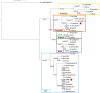
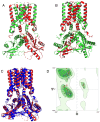
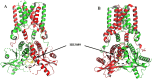
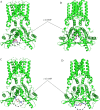
Methods: We structurally characterized the ancestral STING ortholog from Atlantic salmon (Salmo salar) using AlphaFold-guided modeling to identify conserved motifs, including the cyclic dinucleotide (CDN)-binding cleft and phosphorylation regulatory sites. Molecular docking simulations were performed to evaluate the interaction of a validated human STING agonist with salmonid STING. Transcriptomic analyses were conducted in immune tissues and SHK-1 macrophage-like cells infected with Piscirickettsia salmonis to assess gene expression dynamics.
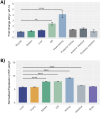
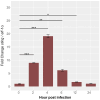
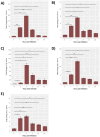
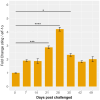
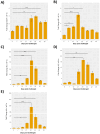
Results: Our models confirmed evolutionary conservation of key STING structural domains. Docking revealed a strong binding affinity between the human agonist and salmonid STING, supporting translational potential. Transcriptomics showed high sting1 expression in immune tissues, rapidly upregulated after infection. In SHK-1 cells, STING1, IFN-α, TNF-α, and IL-1β peaked at 4 hours post-infection (hpi), but this inflammatory burst collapsed by 5 days post-infection (dpi), despite persistent sting1 transcription, indicating functional uncoupling due to immune evasion. In vivo, prolonged DDX41-STING activation was associated with reduced pyroptosis, necroptosis, and inflammatory signaling, reflecting bacterial suppression mechanisms.
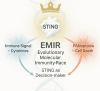
Discussion: This study positions S. salar as a high-resolution model for STING biology and introduces the Evolutionary Molecular Immunity Race (EMIR) framework, where STING orchestrates immune fate across hundreds of millions of years of vertebrate evolution, and over the last ~80 million years within the salmonid lineage.
Keywords: Atlantic salmon-pathogen interaction; STING gene; dynamics activation gene expression; evolutionary perspectives; innate immune response; structural functions.
Copyright © 2025 Yañez, Beltrán, Barrientos, Soto-Rauch, Aguilar, Isla, Flores-Martin, Yañez, Yuivar, Ojeda, Almendras, Bustos and Mancilla.
Conflict of interest statement
The authors PB, MM, YY, and AO were employed by ADL Diagnostic Chile. The authors AY and FA were employed by Greenvolution Spa. Author AY was also employed by KeyBio Solution. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

