Cytidine diphosphate diacylglycerol synthase 2 is a synthetic lethal target in mesenchymal-like cancers
- PMID: 40615674
- PMCID: PMC12283369
- DOI: 10.1038/s41588-025-02221-2
Cytidine diphosphate diacylglycerol synthase 2 is a synthetic lethal target in mesenchymal-like cancers
Abstract
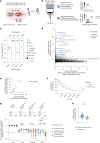
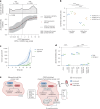
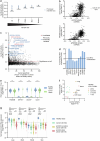
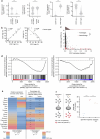
Synthetic lethal interactions (SLIs) based on genomic alterations in cancer have been therapeutically explored. We investigated the SLI space as a function of differential RNA expression in cancer and normal tissue. Computational analyses of functional genomic and gene expression resources uncovered a cancer-specific SLI between the paralogs cytidine diphosphate diacylglycerol synthase 1 (CDS1) and CDS2. The essentiality of CDS2 for cell survival is observed for mesenchymal-like cancers, which have low or absent CDS1 expression and account for roughly half of all cancers. Mechanistically, the CDS1-2 SLI is accompanied by disruption of lipid homeostasis, including accumulation of cholesterol esters and triglycerides, and apoptosis. Genome-wide CRISPR-Cas9 knockout screens in CDS1-negative cancer cells identify no common escape mechanism of death caused by CDS2 ablation, indicating the robustness of the SLI. Synthetic lethality is driven by CDS2 dosage and depends on catalytic activity. Thus, CDS2 may serve as a pharmacologically tractable target in mesenchymal-like cancers.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: T.A. and D.S.P. are named inventors on a patent application related to this work (no. WO2024240955A1). D.S.P. is co-founder, shareholder and advisor of Flindr Therapeutics, which is unrelated to the present study. The other authors declare no competing interests.
Figures











References
-
- Hartwell, L. H., Szankasi, P., Roberts, C. J., Murray, A. W. & Friend, S. H. Integrating genetic approaches into the discovery of anticancer drugs. Science278, 1064–1068 (1997). - PubMed
-
- Venkitaraman, A. R. Cancer susceptibility and the functions of BRCA1 and BRCA2. Cell108, 171–182 (2002). - PubMed
-
- Bryant, H. E. et al. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature434, 913–917 (2005). - PubMed
-
- Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature434, 917–921 (2005). - PubMed
-
- Fong, P. C. et al. Inhibition of poly(ADP-ribose) polymerase in tumors from mutation carriers. N. Engl. J. Med.361, 123–134 (2009). - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Medical

