ARABIDOPSIS HOMOLOG OF TRITHORAX1 impacts lateral root development by epigenetic regulation of targets involved in root system architecture
- PMID: 40624794
- PMCID: PMC12329177
- DOI: 10.1111/nph.70349
ARABIDOPSIS HOMOLOG OF TRITHORAX1 impacts lateral root development by epigenetic regulation of targets involved in root system architecture
Abstract
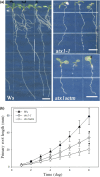
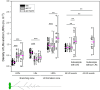
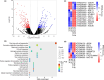
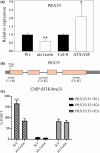
Developmental processes are regulated at multiple levels, including the epigenetic level. Among the epigenetic factors, histone H3 lysine 4 (H3K4) methyltransferases contribute to active transcription of target genes, and here, we explored how the H3K4 methyltransferase ARABIDOPSIS HOMOLOG OF TRITHORAX1 (ATX1) affects Arabidopsis thaliana lateral root (LR) primordium (LRP) morphogenesis. We examined LR development in a loss-of-function null mutant (atx1-1) and a mutant affected in the ATX1 catalytic domain (atx1setm) through bright-field and long-term time-lapse confocal microscopy, transcriptomics, and chromatin immunoprecipitation. LRP morphogenesis in both mutants was severely abnormal, resulting from altered principal growth directions, and was accompanied by extended cell cycle durations and slower transitions between LRP stages. Among the differentially expressed genes downregulated in atx1setm, the most enriched Gene Ontology categories were cell wall organization and H2O2 metabolism, the latter of which included PEROXIDASE35 (PRX35). The LRP morphogenesis abnormalities were similar in prx35 and atx1 mutants. Both the deposition of H3K4me3 at the PRX35 promoter and the PRX35 expression in the atx1setm mutant were significantly reduced. Our results reveal a link between LR development and a redox homeostasis controlled by ATX1 at epigenetic level by maintaining active transcription of PRX35 and thereby impacting root system formation.
Los procesos de desarrollo están regulados a diferentes niveles incluyendo el nivel epigenético. La histona metiltransferasa (H3K4me) es un factor epigenético que contribuye a la transcripción activa de sus genes diana. En este trabajo exploramos, como la H3K4 metiltransferasa ARABIDOPSIS HOMOLOG OF TRITHORAX1 (ATX1) afecta la morfogénesis del primordio de raíz lateral, proceso fundamental para que la raíz lateral se desarrolle adecuadamente. Usando diferentes enfoques, tales como microscopía de campo claro y confocal, experimentos de rastreo celular en tejidos vivos, análisis del transcriptoma e inmunoprecipitación de la cromatina, estudiamos el desarrollo de la raíz lateral en una mutante nula de pérdida de función (atx1‐1) y en una mutante afectada en el dominio catalítico de ATX1 (atx1setm), ambas en planta modelo Arabidopsis thaliana. Encontramos que la morfogénesis del primordio de raíz lateral resultó severamente afectada en las mutantes como consecuencia del cambio en la dirección principal del crecimiento, del ciclo celular alargado y de las transiciones lentas entre los estadios del desarrollo de los primordios. Las categorias de ontología de genes más enriquecidas entre los genes diferencialmente expresados que bajaron su abundancia en la mutante atx1‐1, fueron la organización de la pared celular y el metabolismo del peróxido de hidrogeno, dentro de esta última categoría se encontró a la PEROXIDASE35 (PRX35). Las anomalías en la morfogénesis de los primordios de raíz lateral en las mutantes prx35 fueron similares a las anomalías encontradas en atx1. Evidenciamos que en la mutante atx1setm tanto la deposición de la marca H3K4me3 en el promotor de PRX35 como su expresión se disminuyeron significativamente, lo que sugiere que ATX1 regula a PRX35 y ambos genes son esenciales para el desarrollo de la raíz en plantas. Este trabajo revelo una asociación entre el control del desarrollo de la raíz lateral a nivel epigenético y la homeostasis del estado redox, involucrado en la morfogénesis del primordio de la raíz lateral. Esta asociación impacta el control de la arquitectura del sistema radical en plantas.
Keywords: cell cycle; epigenetic regulation; lateral roots; live imaging; methyltransferase activity; morphogenesis; reactive oxygen species; transcriptomics.
© 2025 The Author(s). New Phytologist © 2025 New Phytologist Foundation.
Conflict of interest statement
None declared.
Figures



Similar articles
-
Arabidopsis homolog of trithorax1 (ATX1) is required for cell production, patterning, and morphogenesis in root development.J Exp Bot. 2014 Dec;65(22):6373-84. doi: 10.1093/jxb/eru355. Epub 2014 Sep 9. J Exp Bot. 2014. PMID: 25205583 Free PMC article.
-
RGIs-mediated root apical meristem development is essential for root hydrotropic response in Arabidopsis thaliana.Plant J. 2025 Jun;122(6):e70273. doi: 10.1111/tpj.70273. Plant J. 2025. PMID: 40561222
-
Histone methyltransferase ATX1 dynamically regulates fiber secondary cell wall biosynthesis in Arabidopsis inflorescence stem.Nucleic Acids Res. 2021 Jan 11;49(1):190-205. doi: 10.1093/nar/gkaa1191. Nucleic Acids Res. 2021. PMID: 33332564 Free PMC article.
-
Stage-specific DNA methylation dynamics in mammalian heart development.Epigenomics. 2025 Apr;17(5):359-371. doi: 10.1080/17501911.2025.2467024. Epub 2025 Feb 21. Epigenomics. 2025. PMID: 39980349 Review.
-
ATX1/AtCOMPASS and the H3K4me3 marks: how do they activate Arabidopsis genes?Curr Opin Plant Biol. 2014 Oct;21:75-82. doi: 10.1016/j.pbi.2014.07.004. Epub 2014 Jul 19. Curr Opin Plant Biol. 2014. PMID: 25047977 Review.
References
-
- Alvarez‐ Buylla ER, Liljergen J, Pelaz S, Gold SE, Burgeff C, Ditta GS, Vergara‐Silva F, Yanofsky MF. 2000a. MADS‐box gene evolution beyond flowers: expression in pollen, endosperm, guard cells, roots and trichomes. The Plant Journal 24: 457–466. - PubMed
-
- Alvarez‐Venegas R, Pien S, Sadder M, Witmer X, Grossniklaus U, Avramova Z. 2003. ATX‐1, an Arabidopsis homolog of trithorax, activates flower homeotic genes. Current Biology 13: 627–637. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

