Single-cell analysis links DCUN1D5 to immune remodeling and cisplatin resistance in recurrent osteosarcoma
- PMID: 40629059
- PMCID: PMC12238620
- DOI: 10.1038/s42003-025-08409-w
Single-cell analysis links DCUN1D5 to immune remodeling and cisplatin resistance in recurrent osteosarcoma
Abstract
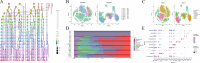
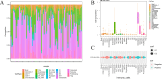
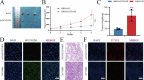
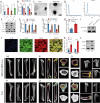
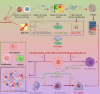
Cisplatin is the primary chemotherapeutic agent for osteosarcoma. However, a significant proportion of patients develop resistance post-treatment, leading to disease recurrence and presenting profound clinical challenges. To understand the mechanisms underlying osteosarcoma recurrence and cisplatin resistance, particularly from the tumor microenvironment perspective, we consolidated numerous single-cell RNA sequencing datasets, offering an encompassing insight into the osteosarcoma microenvironment. When juxtaposing scRNA-seq with bulk RNA-seq data, we observed a strong correlation between high DCUN1D5 expression in osteosarcoma and patient survival. This gene amplifies osteosarcoma's anti-apoptotic, invasive, stem-cell-like traits and PI3K/AKT/GSK3β pathway phosphorylation and fosters cisplatin resistance. Subsequent research revealed that cisplatin-resistant osteosarcoma cells excrete DCUN1D5-rich exosomes, facilitating the maturation of osteoclast precursors. Excessive osteoclast activity is a pivotal contributor to osteosarcoma recurrence and resistance. Given these insights, DCUN1D5 is a promising therapeutic target for osteosarcoma recurrence and drug resistance.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests. The authors declare no competing interests. Figure 6I, Figure S13A and graphic abstract, was created with BioRender software ( https://biorender.com/ , accessed on 3 March 2023). Ethical approval: All animal care and experimental procedures were conducted in accordance with the guidelines and were approved by the Ethics Committee of the Department of Laboratory Animal Science, Central South University (Approval Number: CSU-2022-0664; Approval Date: October 10, 2022). We have complied with all relevant ethical regulations for animal use.
Figures






Similar articles
-
A Dual Approach with Organoid and CRISPR Screening Reveals ERCC6 as a Determinant of Cisplatin Resistance in Osteosarcoma.Adv Sci (Weinh). 2025 Jul;12(28):e2500632. doi: 10.1002/advs.202500632. Epub 2025 Jun 6. Adv Sci (Weinh). 2025. PMID: 40476445 Free PMC article.
-
Formononetin enhances cisplatin chemotherapy sensitivity in osteosarcoma by inducing ferroptosis and reconstructing the immune microenvironment.Phytomedicine. 2025 Sep;145:156960. doi: 10.1016/j.phymed.2025.156960. Epub 2025 Jun 9. Phytomedicine. 2025. PMID: 40543234
-
The EphA2 Receptor Regulates Invasiveness and Drug Sensitivity in Canine and Human Osteosarcoma Cells.Cells. 2024 Jul 16;13(14):1201. doi: 10.3390/cells13141201. Cells. 2024. PMID: 39056783 Free PMC article.
-
Methotrexate for high-grade osteosarcoma in children and young adults.Cochrane Database Syst Rev. 2011 May 11;2011(5):CD006325. doi: 10.1002/14651858.CD006325.pub3. Cochrane Database Syst Rev. 2011. PMID: 21563152 Free PMC article.
-
Methotrexate for high-grade osteosarcoma in children and young adults.Cochrane Database Syst Rev. 2009 Jan 21;(1):CD006325. doi: 10.1002/14651858.CD006325.pub2. Cochrane Database Syst Rev. 2009. Update in: Cochrane Database Syst Rev. 2011 May 11;(5):CD006325. doi: 10.1002/14651858.CD006325.pub3. PMID: 19160278 Updated.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

