Exploring the molecular mechanism of Polygonum multiflorum in treating androgenic alopecia based on the methods of bioinformatics and molecular docking
- PMID: 40629596
- PMCID: PMC12237353
- DOI: 10.1097/MD.0000000000043025
Exploring the molecular mechanism of Polygonum multiflorum in treating androgenic alopecia based on the methods of bioinformatics and molecular docking
Abstract
This study aims to research and discuss the molecular mechanism of Polygonum multiflorum (PM) to treat androgenic alopecia (APA). The target network and protein interaction network of APA of PM were established using traditional Chinese medicine molecular mechanism bioinformatics databases and disease gene databases. The core subnetworks were screened based on topological analysis, and Gene Ontology and Kyoto Encyclopedia of Genes and Genomes enrichment analyses were performed, molecular docking of the 10 genes with the highest connectivity. According to the intersection of active components and potential targets of PM, 1325 common target factors were obtained. The core subnetwork composed of 66 genes was screened by topological analysis. The results of Kyoto Encyclopedia of Genes and Genomes enrichment showed that PM mainly exerts biological functions by regulating inflammatory response, oxidation stress, apoptosis, autophagy, and other pathways. The result of molecular docking shows that the binding energies of HRAS, PIK3CA, PIK3CB, and RHOA -resveratrol, -polygonin, -chrysophanol, -rheum emodin, and -rhein were all <-9 kcal/mol. PM's mechanism in treating APA may regulate the expression of PIK3CA and PIK3CB through the emodin-mediated phosphoinositide 3-kinase/protein kinase B pathway to regulate cell apoptosis. In addition, some processes, such as inhibiting inflammation, oxidative stress, and inducing melanin production, are also involved. PM has application value in treating APA and can be popularized after testing.
Keywords: Polygonum multiflorum; androgenic alopecia; bioinformatic; molecular docking; molecular mechanism.
Copyright © 2025 the Author(s). Published by Wolters Kluwer Health, Inc.
Conflict of interest statement
The authors have no conflicts of interest to disclose.
Figures
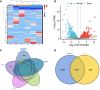



References
-
- Rajabi F, Drake LA, Senna MM, Rezaei N. Alopecia areata: a review of disease pathogenesis. Br J Dermatol. 2018;179:1033–48. - PubMed
-
- Chen D, Yang X, Liu X, et al. Efficacy comparison of monotherapies and combination therapies for androgenetic alopecia: a Bayesian network meta-analysis. Dermatol Ther. 2022;35:e15262. - PubMed
-
- Dhurat RS, Daruwalla SB. Androgenetic alopecia: update on etiology. Dermatol Rev. 2021;2:115–21.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

