Molecular epidemiology and clinical characteristics of carbapenem-resistant Klebsiella pneumoniae bloodstream and pneumonia isolates
- PMID: 40631745
- PMCID: PMC12323367
- DOI: 10.1128/spectrum.00631-25
Molecular epidemiology and clinical characteristics of carbapenem-resistant Klebsiella pneumoniae bloodstream and pneumonia isolates
Abstract
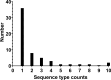
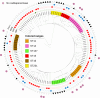
Appropriate antibiotic therapy in patients with carbapenem-resistant Klebsiella pneumoniae (CRKp) infections often requires knowledge of resistance mechanisms. Therefore, we aimed to define the clonal diversity, mechanisms of carbapenem resistance, potential for carbapenem resistance transmissibility, virulence gene repertoire, and potential clinical impact of these genetic elements in patients with CRKp bloodstream infection and/or pneumonia, which are associated with high mortality. Clinical data and CRKp isolates at a site in Singapore and in the United States were examined. Whole-genome sequencing and bioinformatic analysis of CRKp isolates were performed. In total, 125 patients (n = 118 from Singapore; n = 7 from the United States) were included. Clonal diversity was high as 58 sequence types were identified. Carbapenemases were present in 109 (87%) of the CRKp isolates; KPC-, OXA-, and NDM-type enzymes were identified in significant quantity. The majority (87%) of carbapenemases were encoded on conjugative or mobilizable plasmids and thus potentially transmissible to other strains. Of the 16 CRKp isolates without a carbapenemase, the predominant predicted mechanism of carbapenem resistance was extended-spectrum β-lactamases along with truncation of porins ompK35 and/or ompK36. Patients with carbapenemase-producing CRKp strains had increased in-hospital mortality. Recurrent CRKp infections were primarily due to either the same CRKp genomic group or similar carbapenemase-encoding conjugative plasmids within a different K. pneumoniae strain. Our findings suggest that in CRKp isolates causing serious infections, carbapenem resistance arises through diverse mechanisms, is easily transmissible between isolates, and impacts patient outcomes. These findings highlight the importance of diagnostic tools to identify mechanisms of carbapenem resistance in the clinical setting.IMPORTANCETo design more effective therapies and better understand treatment failure in patients with carbapenem-resistant Klebsiella pneumoniae (CRKp) infections, we must identify the mechanisms of carbapenem resistance and their impact on patient outcomes. However, prior studies are often limited by the inclusion of CRKp that are colonizers or from non-severe infections, a focus on a single general mechanism (i.e., presence of carbapenemases), lack of whole-genome sequencing to fully characterize the underlying genetic architecture of CRKp strains, and no analyses of associations between resistance mechanisms and clinical outcomes. Here, we attempt to address some of these gaps by sequencing a large set of invasive CRKp isolates to characterize the underlying clonal diversity, mechanisms of carbapenem resistance, potential carbapenemase transmissibility, virulence gene repertoire, and the possible impact of these genetic elements on patient mortality.
Keywords: Klebsiella pneumoniae; antibiotic resistance; carbapenem.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Chen Y, Chen Y, Liu P, Guo P, Wu Z, Peng Y, Deng J, Kong Y, Cui Y, Liao K, Huang B. 2022. Risk factors and mortality for elderly patients with bloodstream infection of carbapenem resistance Klebsiella pneumoniae: a 10-year longitudinal study. BMC Geriatr 22:573. doi: 10.1186/s12877-022-03275-1 - DOI - PMC - PubMed
-
- Zhen X, Stålsby Lundborg C, Sun X, Gu S, Dong H. 2020. Clinical and economic burden of carbapenem-resistant infection or colonization caused by Klebsiella pneumoniae, Pseudomonas aeruginosa, Acinetobacter baumannii: a multicenter study in China. Antibiotics (Basel) 9:514. doi: 10.3390/antibiotics9080514 - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

