Identification and validation of epithelial‑mesenchymal transition‑related genes for diabetic nephropathy by WGCNA and machine learning
- PMID: 40641111
- PMCID: PMC12272152
- DOI: 10.3892/mmr.2025.13614
Identification and validation of epithelial‑mesenchymal transition‑related genes for diabetic nephropathy by WGCNA and machine learning
Abstract
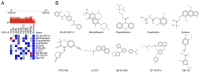
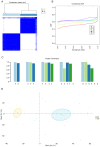
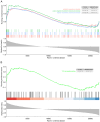
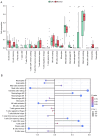
Diabetic nephropathy (DN) is the main cause of end‑stage renal disease, with epithelial‑mesenchymal transition (EMT) serving a key role in its initiation and progression. Nevertheless, the precise mechanisms involved remain unidentified. The present study aimed to identify the involvement of EMT‑related genes in the advancement of DN. Using the Gene Expression Omnibus database and the dbEMT 2.0 database, EMT‑related differentially expressed genes (DEGs) associated with DN were identified. Key EMT‑related genes were subjected to weighted gene co‑expression network analysis, machine learning and protein‑protein interaction network analyses and validated against validation datasets from GEO database. Receiver operating characteristic analysis was used to assess the diagnostic performance of these hub genes. To delve into their cellular clustering in DN, single‑nucleus RNA sequencing was conducted using the Kidney Integrative Transcriptomics database. Additionally, the CIBERSORT algorithm was used to determine the proportion of immune cell infiltration in DN samples. Reverse transcription‑quantitative PCR (RT‑qPCR) was used to assess the mRNA expression of fibronectin 1 (FN1) in the kidney of mice and patients with DN. After silencing FN1, the expression changes of EMT markers (E‑cadherin and vimentin) were detected by RT‑qPCR. FN1 was upregulated in DN, demonstrating good diagnostic performance according to ROC analysis. FN1 was associated with infiltration of immune cells. RT‑qPCR confirmed the increased expression of FN1 in the kidney of mice with DN and in the renal biopsy samples of patients with DN. After silencing FN1, the expression of E‑cadherin was upregulated, while the expression of vimentin was downregulated, indicating that EMT was inhibited. The present study identified FN1 as a diagnostic marker for DN. FN1 may serve key roles in the initiation and progression of DN by participating in EMT and upregulating various types of immune cells.
Keywords: FN1; clustering analysis; diabetic nephropathy; immune cell infiltration; machine learning; weighted gene co‑expression network analysis.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures





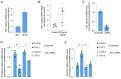
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

