Array-based polymer-phage biosensors for detection and differentiation of bacteria
- PMID: 40641937
- PMCID: PMC12235245
- DOI: 10.1039/d5sd00069f
Array-based polymer-phage biosensors for detection and differentiation of bacteria
Abstract
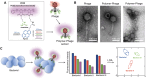
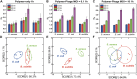
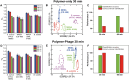
Pathogenic bacteria, such as methicillin-resistant Staphylococcus aureus (MRSA), pose significant challenges to public health due to their resistance to conventional antibiotics. Early and accurate identification of bacterial species and discrimination of their strains is critical for guiding effective treatments and infection control. In this study, we develop a polymer-phage sensor platform that integrates polymer-based fluorescence sensing with phage-host specificity for bacterial identification. The sensor successfully differentiates three bacterial species (S. aureus, E. coli, and B. subtilis) and closely related strains of S. aureus (methicillin-sensitive Staphylococcus aureus (MSSA) and MRSA) with high classification accuracy (94-100%) and correct unknown identification rates (94-100%) under optimized conditions. By leveraging phage-host interactions and polymer binding properties, the polymer-phage sensor overcomes the limitations of traditional "lock-and-key" biosensors, offering enhanced specificity and reliability. This platform's rapid response time and adaptability make it a promising tool for clinical diagnostics and public health applications, particularly in combating antibiotic-resistant bacteria.
This journal is © The Royal Society of Chemistry.
Conflict of interest statement
There are no conflicts to declare.
Figures
References
-
- WHO, WHO bacterial priority pathogens list, 2024, 2024
Grants and funding
LinkOut - more resources
Full Text Sources
