The STING pathway drives noninflammatory neurodegeneration in NGLY1 deficiency
- PMID: 40644312
- PMCID: PMC12249164
- DOI: 10.1084/jem.20242296
The STING pathway drives noninflammatory neurodegeneration in NGLY1 deficiency
Abstract
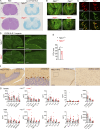
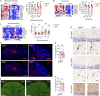
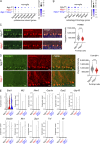
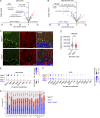
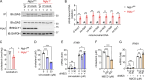
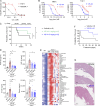
The STING pathway is increasingly recognized as a key regulator of neuroinflammation in neurodegenerative disease, but its role in noninflammatory conditions remains unclear. We generated a postnatal inducible whole-body Ngly1 knockout mouse (iNgly1-/-) to model NGLY1 deficiency, an early-onset neurodegenerative disorder. iNgly1-/- mice exhibit progressive motor deficits, Purkinje cell loss, and shortened lifespan without evidence of gliosis or immune activation. Cell type-specific deletion of Ngly1 in Purkinje cells or microglia failed to induce disease, suggesting multiple cell-intrinsic and cell-extrinsic signals are required. Genetic ablation of Sting1 in iNgly1-/- mice rescues Purkinje cell loss, improves motor function, and extends lifespan. Single-nucleus RNA sequencing reveals proteostasis disruption in Purkinje cells, altered cerebellar granule cell subpopulations, and STING-dependent suppression of cholesterol biosynthesis in glia. Pharmacological inhibition of STING with an orally bioactive antagonist, VS-X4, significantly mitigates neuropathology and motor disease. These findings identify STING as a key mediator of neuropathology in NGLY1 deficiency and implicate a role of STING in noninflammatory neurological disease.
© 2025 Yang et al.
Conflict of interest statement
Disclosures: W.F. Mueller reported personal fees from Grace Science, LLC, outside the submitted work; in addition, W.F. Mueller had a patent to US20240218396A1 issued. K. Lee is an employee of the Grace Science Foundation, a nonprofit foundation that provided partial support for this study. No other disclosures were reported.
Figures






References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

