P4-ATPases control phosphoinositide membrane asymmetry and neomycin resistance
- PMID: 40646185
- PMCID: PMC12270916
- DOI: 10.1038/s41556-025-01692-z
P4-ATPases control phosphoinositide membrane asymmetry and neomycin resistance
Abstract
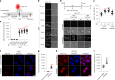
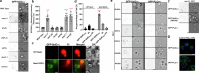
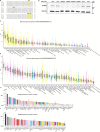
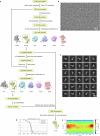
The aminoglycoside antibiotic neomycin has robust antibacterial properties, yet its clinical utility is curtailed by its nephrotoxicity and ototoxicity. The mechanism by which the polycationic neomycin enters specific eukaryotic cell types remains poorly understood. In budding yeast, NEO1 is required for neomycin resistance and encodes a phospholipid flippase that establishes membrane asymmetry. Here we show that mutations altering Neo1 substrate recognition cause neomycin hypersensitivity by exposing phosphatidylinositol-4-phosphate (PI4P) in the plasma membrane extracellular leaflet. Cryogenic electron microscopy reveals PI4P binding to Neo1 within the substrate translocation pathway. PI4P enters the lumen of the endoplasmic reticulum and is flipped by Neo1 at the Golgi to prevent PI4P secretion to the cell surface. Deficiency of the orthologous ATP9A in human cells also causes exposure of PI4P and neomycin sensitivity. These findings unveil conserved mechanisms of aminoglycoside sensitivity and phosphoinositide homoeostasis, with important implications for signalling by extracellular phosphoinositides.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures









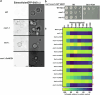
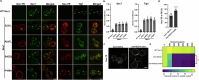
Update of
-
P4-ATPase control over phosphoinositide membrane asymmetry and neomycin resistance.bioRxiv [Preprint]. 2025 Mar 3:2025.03.03.641220. doi: 10.1101/2025.03.03.641220. bioRxiv. 2025. Update in: Nat Cell Biol. 2025 Jul;27(7):1114-1124. doi: 10.1038/s41556-025-01692-z. PMID: 40093091 Free PMC article. Updated. Preprint.
References
-
- Waksman, S. A. & Lechevalier, H. A. Neomycin, a new antibiotic active against streptomycin-resistant bacteria, including tuberculosis organisms. Science109, 305–307 (1949). - PubMed
-
- Robson, J. M. & Sullivan, F. M. Antituberculosis drugs. Pharmacol. Rev.15, 169–223 (1963). - PubMed
-
- Davies, J. & Davis, B. D. Misreading of ribonucleic acid code words induced by aminoglycoside antibiotics. The effect of drug concentration. J. Biol. Chem.243, 3312–3316 (1968). - PubMed
-
- Davies, J., Gorini, L. & Davis, B. D. Misreading of RNA codewords induced by aminoglycoside antibiotics. Mol. Pharmacol.1, 93–106 (1965). - PubMed
-
- Andre, T. Studies on the distribution of tritium-labelled dihydrostreptomycin and tetracycline in the body. Acta Radiol. Suppl.142, 1–89 (1956). - PubMed
MeSH terms
Substances
Grants and funding
- R01CA231466/U.S. Department of Health & Human Services | NIH | National Institute of General Medical Sciences (NIGMS)
- R35GM144123/U.S. Department of Health & Human Services | NIH | National Institute of General Medical Sciences (NIGMS)
- S10 OD021630/OD/NIH HHS/United States
- R01 CA231466/CA/NCI NIH HHS/United States
- R35 GM144123/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources

