Phylogenetic and Structural Insights into Melatonin Receptors in Plants: Case Study in Capsicum chinense Jacq
- PMID: 40647961
- PMCID: PMC12252195
- DOI: 10.3390/plants14131952
Phylogenetic and Structural Insights into Melatonin Receptors in Plants: Case Study in Capsicum chinense Jacq
Abstract
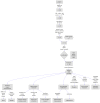
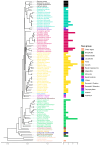
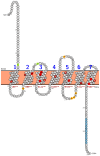
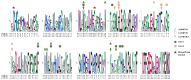
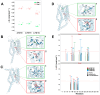
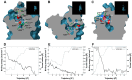
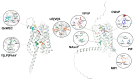
Recently, it has been proposed that plant melatonin receptors belong to the superfamily of G protein-coupled receptors (GPCRs). However, a detailed description of the phylogeny, protein structure, and binding properties of melatonin, which is still lacking, can help determine the signaling and function of this compound. Melatonin receptor homologs (PMTRs) were identified in 90 Viridiplantae sensu lato proteomes using profile Hidden Markov Models (HMM), which yielded 174 receptors across 87 species. Phylogenetic analysis revealed an expansion of PMTR sequences in angiosperms, which were grouped into three clades. Docking studies uncovered a conserved internal melatonin-binding site in PMTRs, which was analogous to the site in human MT1 receptors. Binding affinity simulations indicated this internal site exhibits stronger melatonin binding compared to a previously reported superficial pocket. Ligand-receptor interaction analysis and alanine scanning highlighted a major role of hydrophobic interactions, with hydrogen bonds contributing predominantly at the internal site, while non-interacting charged residues stabilize the binding pocket. Tunnel and ligand transport simulations suggested melatonin moves favorably through the internal cavity to access the binding site. Also, we presented for the first time details of these pockets in a non-model species, Capsicum chinense. Taken together, the structural analyses presented here illustrate opportunities and theoretical evidence for performing structure-function studies via mutations in specific residues within the proposed new melatonin-binding site in PMTRs, shedding light on their role in plant melatonin signaling.
Keywords: Arabidopsis; Capsicum; G protein-coupled receptor; HMM; melatonin; molecular docking; phylogeny; phytomelatonin receptor PMTR.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures



Similar articles
-
The Black Book of Psychotropic Dosing and Monitoring.Psychopharmacol Bull. 2024 Jul 8;54(3):8-59. Psychopharmacol Bull. 2024. PMID: 38993656 Free PMC article. Review.
-
[Volume and health outcomes: evidence from systematic reviews and from evaluation of Italian hospital data].Epidemiol Prev. 2013 Mar-Jun;37(2-3 Suppl 2):1-100. Epidemiol Prev. 2013. PMID: 23851286 Italian.
-
Structural basis of ligand recognition at the human MT1 melatonin receptor.Nature. 2019 May;569(7755):284-288. doi: 10.1038/s41586-019-1141-3. Epub 2019 Apr 24. Nature. 2019. PMID: 31019306 Free PMC article.
-
Short-Term Memory Impairment.2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 31424720 Free Books & Documents.
-
The Lived Experience of Autistic Adults in Employment: A Systematic Search and Synthesis.Autism Adulthood. 2024 Dec 2;6(4):495-509. doi: 10.1089/aut.2022.0114. eCollection 2024 Dec. Autism Adulthood. 2024. PMID: 40018061 Review.
References
Grants and funding
LinkOut - more resources
Full Text Sources

