Molecular Mechanisms Underlying Defense Responses of Potato (Solanum tuberosum L.) to Environmental Stress and CRISPR/Cas-Mediated Engineering of Stress Tolerance
- PMID: 40647992
- PMCID: PMC12252150
- DOI: 10.3390/plants14131983
Molecular Mechanisms Underlying Defense Responses of Potato (Solanum tuberosum L.) to Environmental Stress and CRISPR/Cas-Mediated Engineering of Stress Tolerance
Abstract
Environmental stresses, such as drought, salinity, and pathogen attacks, significantly affect potato growth, development, and yield by disrupting key physiological and biochemical processes. Plant responses to these stresses are mediated by changes in gene expression, transcriptional regulation, and the activity of various functional proteins, all of which contribute to the molecular mechanisms of stress tolerance. Genome editing using the CRISPR/Cas9 system has been effectively used to enhance the resistance of potato to environmental stresses and to improve its nutritional value. This article provides a comprehensive review of recent studies retrieved from academic databases focusing on the effects of various environmental stressors on potato growth, yield, and postharvest storage. It also examines the influence of these stresses on the production of secondary metabolites and their associated molecular pathways. Finally, the review highlights advancements in the application of CRISPR/Cas-based genome editing technologies between 2021 and 2025 to improve stress tolerance and nutritional traits in potato plants.
Keywords: CRISPR/Cas; Solanum tuberosum; defense mechanisms; environmental stresses; secondary metabolites.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
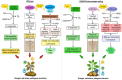

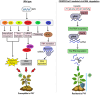
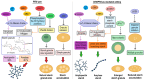
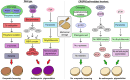
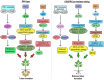
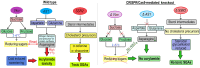
References
-
- Nasir M.W., Toth Z. Effect of Drought Stress on Potato Production: A Review. Agronomy. 2022;12:635. doi: 10.3390/agronomy12030635. - DOI
-
- Demirel U. Chapter 4—Environmental requirements of potato and abiotic stress factors. In: Çalişkan M.E., Bakhsh A., Jabran K., editors. Potato Production Worldwide. Academic Press; Cambridge, MA, USA: 2023. pp. 71–86.
-
- Gibson S., Kurilich A.C. The nutritional value of potatoes and potato products in the UK diet. Nutr. Bull. 2013;38:389–399. doi: 10.1111/nbu.12057. - DOI
-
- Haverkort A., Struik P. Yield levels of potato crops: Recent achievements and future prospects. Field Crops Res. 2015;182:76–85. doi: 10.1016/j.fcr.2015.06.002. - DOI
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources

