Phenotypic Plasticity and Androgen Receptor Bypass Drive Cross-Resistance to Apalutamide in Castration-Resistant Prostate Cancer Cell Models
- PMID: 40649715
- PMCID: PMC12250327
- DOI: 10.3390/ijms26135939
Phenotypic Plasticity and Androgen Receptor Bypass Drive Cross-Resistance to Apalutamide in Castration-Resistant Prostate Cancer Cell Models
Abstract
The treatment of choice for prostate cancer is androgen deprivation (ADT) and novel hormonal agents such as Abiraterone, Enzalutamide, or Apalutamide. Initially, this therapy is highly effective, but a significant challenge arises as most patients eventually develop resistance, resulting in castration-resistant prostate cancer (CRPC). Furthermore, the sequential use of these drugs can lead to cross-resistance, diminishing their efficacy. Tumor heterogeneity plays a pivotal role in the development of resistance to different treatments. This study utilized cellular models of CRPC to assess the response to Apalutamide when it was administered as a second- or third-line treatment. Functional and genetic analyses were conducted in various CRPC cell models exposed to Apalutamide. These analyses included real-time cell monitoring assays, flow cytometry, clonogenicity assays, and RT-qPCR. CRPC cell models were capable of continued proliferation, maintained cell cycle profiles similar to those of untreated cells, and retained their clonogenic potential. Cross-resistance to Apalutamide in models of ADT, ADT plus Enzalutamide, or Abiraterone resistance did not correlate with the expression levels of AR-V7 and AR-V9 variants. Gene expression analysis of resistant prostate cancer cell lines revealed that treatment with Apalutamide induced the emergence of more aggressive phenotypes, including cancer stem cells or neuroendocrine differentiation profiles. Most CRPC cell models developed cross-resistance to Apalutamide and were able to proliferate and retain their clonogenic capability. Apalutamide resistance was not linked to the expression of AR-V7 or AR-V9 variants but was instead associated to bypass of AR signaling pathway and the emergence of more aggressive expression profiles.
Keywords: Abiraterone; Apalutamide; Enzalutamide; androgen deprivation therapy (ADT); castration-resistant prostate cancer (CRPC); prostate cancer.
Conflict of interest statement
The authors declare no conflicts of interest.
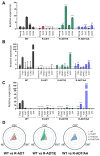
Figures


Similar articles
-
Loss of AR-regulated AFF3 contributes to prostate cancer progression and reduces ferroptosis sensitivity by downregulating ACSL4 based on single-cell sequencing analysis.Apoptosis. 2024 Oct;29(9-10):1679-1695. doi: 10.1007/s10495-024-01941-w. Epub 2024 Mar 13. Apoptosis. 2024. PMID: 38478171
-
Targeted Delivery of AR-V7 siRNA with Bivalent PSMA Aptamers Effectively Suppresses the Growth of Enzalutamide-Resistant Prostate Cancer.Mol Pharm. 2024 Nov 4;21(11):5749-5760. doi: 10.1021/acs.molpharmaceut.4c00743. Epub 2024 Oct 10. Mol Pharm. 2024. PMID: 39388218
-
Matching-Adjusted Indirect Comparison of the Efficacy of Apalutamide and Enzalutamide with ADT in the Treatment of Non-Metastatic Castration-Resistant Prostate Cancer.Adv Ther. 2020 Jan;37(1):501-511. doi: 10.1007/s12325-019-01156-5. Epub 2019 Dec 7. Adv Ther. 2020. PMID: 31813086 Free PMC article.
-
EAU guidelines on prostate cancer. Part II: Treatment of advanced, relapsing, and castration-resistant prostate cancer.Eur Urol. 2014 Feb;65(2):467-79. doi: 10.1016/j.eururo.2013.11.002. Epub 2013 Nov 12. Eur Urol. 2014. PMID: 24321502
-
Comparative efficacy of second-generation androgen receptor inhibitors for treating prostate cancer: A systematic review and network meta-analysis.Front Endocrinol (Lausanne). 2023 Mar 9;14:1134719. doi: 10.3389/fendo.2023.1134719. eCollection 2023. Front Endocrinol (Lausanne). 2023. PMID: 36967752 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

