This is a preprint.
Decoding sexual dimorphism of the sex-shared nervous system at single-neuron resolution
- PMID: 40654890
- PMCID: PMC12247729
- DOI: 10.1101/2024.12.27.630541
Decoding sexual dimorphism of the sex-shared nervous system at single-neuron resolution
Update in
-
Decoding sexual dimorphism of the sex-shared nervous system at single-neuron resolution.Sci Adv. 2025 Jul 11;11(28):eadv9106. doi: 10.1126/sciadv.adv9106. Epub 2025 Jul 11. Sci Adv. 2025. PMID: 40644535 Free PMC article.
Abstract
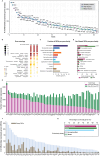
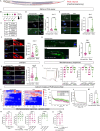
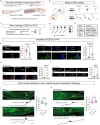
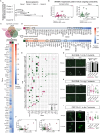
Sex-specific behaviors are often attributed to differences in neuronal wiring and molecular composition, yet how genetic sex shapes the molecular architecture of the nervous system at the individual neuron level remains unclear. Here, we use single-cell RNA sequencing to profile the transcriptome of sex-shared neurons in adult Caenorhabditis elegans males and hermaphrodites. We uncover widespread molecular dimorphism across the nervous system, including in previously unrecognized neuron-types such as the touch receptors. Neuropeptides and signaling-related genes exhibit strong sex-biased expression, particularly in males, reinforcing the notion that neuropeptides are crucial for diversifying connectome outputs. Despite these differences, neurotransmitter identities remain largely conserved, indicating that functional dimorphism arises through modulatory, not identity-defining, changes. We show that sex-biased expression of neurotransmitter-related genes correlates with bias in outgoing synaptic connectivity and identify regulatory candidates for synaptic wiring, including both shared and sex-specific genes. This dataset provides a molecular framework for understanding how subtle regulatory differences tune conserved circuits to drive sex-specific behaviors.
Conflict of interest statement
Competing interests The authors declare no competing interests.
Figures



References
-
- Gater R., Tansella M., Korten A., Tiemens B. G., Mavreas V. G., Olatawura M. O., Sex differences in the prevalence and detection of depressive and anxiety disorders in general health care settings: report from the World Health Organization Collaborative Study on Psychological Problems in General Health Care. Arch Gen Psychiatry 55, 405–413 (1998). - PubMed
-
- Aleman A., Kahn R. S., Selten J. P., Sex differences in the risk of schizophrenia: evidence from meta-analysis. Arch Gen Psychiatry 60, 565–571 (2003). - PubMed
-
- Fang S., Luo Z., Wei Z., Qin Y., Zheng J., Zhang H., Jin J., Li J., Miao C., Yang S., Li Y., Liang Z., Yu X. D., Zhang X. M., Xiong W., Zhu H., Gan W. B., Huang L., Li B., Sexually dimorphic control of affective state processing and empathic behaviors. Neuron 112, 1498–1517.e8 (2024). - PubMed
-
- Chiu H., Hoopfer E. D., Coughlan M. L., Pavlou H. J., Goodwin S. F., Anderson D. J., Erratum: A circuit logic for sexually shared and dimorphic aggressive behaviors in Drosophila (Cell (2021) 184(2) (507–520.e16), (S0092867420316202), ( 10.1016/j.cell.2020.11.048)). Cell 184, 847 (2021). - DOI - PMC - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
