PAXX/Ku interaction is rate limiting for repair of double-strand DNA breaks requiring end processing
- PMID: 40659092
- PMCID: PMC12357295
- DOI: 10.1016/j.jbc.2025.110481
PAXX/Ku interaction is rate limiting for repair of double-strand DNA breaks requiring end processing
Abstract
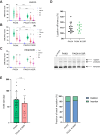
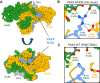
In mammalian cells, DNA double strand breaks (DSBs) are primarily repaired via classical non-homologous end joining (c-NHEJ)-one of the most essential DNA repair pathways. As NHEJ does not utilize a template, this type of repair is the default mechanism for eliminating DSBs occurring in non-cycling cells. NHEJ is a crucial process in mammals, and defects of this repair pathway often result in immunological impairment owing to failure of somatic recombination in lymphocytes and improper neuronal biogenesis. The NHEJ machinery assembles in a stepwise process at DSBs and proceeds via several key repair phases including break recognition, mediated by Ku proteins (Ku70/80 heterodimer binding to DNA), DNA ends processing, and finally DNA ligation. DNA end-bound Ku recruits the large kinase protein DNA-PKcs, and downstream repair-facilitating components such as PAXX, XLF and XRCC4/Ligase IV complex that together facilitate the repair reaction. Processing of DNA breaks can require both nucleotide removal and incorporation, involving a plethora of enzymes such as nucleases and polymerases. It is currently not known which step, if any, limits the completion of the repair process. Here, we describe a single conserved amino acid substitution in PAXX protein Ku70/80 contact interface that dramatically stabilizes the repair complex. This mutation leads to co-dependent mislocalization of PAXX and Ku to the nucleoli. Surprisingly, this novel PAXX gain-of-function mutation accelerates NHEJ repair but only of DSBs that require end processing such as radiation-induced DSBs. Thus, in mammalian NHEJ, the repair complex stability is rate-limiting for the overall repair reaction of DSBs.
Keywords: DNA damage; DNA damage response; DNA repair; DSB; NHEJ; PAXX; double strand break; molecular biology; non-homologous end joining; nucleolus.
Copyright © 2025 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest The authors declare that they have no conflicts of interest with the contents of this article.
Figures


References
-
- Rulten S.L., Fisher A.E., Robert I., Zuma M.C., Rouleau M., Ju L., et al. PARP-3 and APLF function together to accelerate nonhomologous end-joining. Mol. Cell. 2011;41:33–45. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

