Paradoxical mTORC1-Dependent microRNA-mediated Translation Repression in the Nucleus Accumbens of Male Mice Consuming Alcohol Attenuates Glycolysis
- PMID: 40659615
- PMCID: PMC12259992
- DOI: 10.1038/s41467-025-60337-9
Paradoxical mTORC1-Dependent microRNA-mediated Translation Repression in the Nucleus Accumbens of Male Mice Consuming Alcohol Attenuates Glycolysis
Abstract
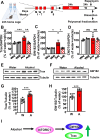
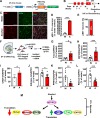
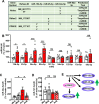
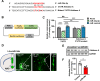
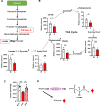
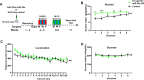
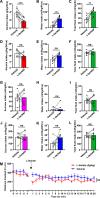
mTORC1 promotes protein translation, learning and memory, and neuroadaptations that underlie alcohol use disorder (AUD). The mechanisms underlying alcohol-mediated mTORC1-dependent neuroadaptations that drive AUD are not well understood. We report that activation of mTORC1 in the nucleus accumbens (NAc) D1 neurons of male mice consuming alcohol results in paradoxical mTORC1-dependent repression of translation of transcripts, including Aldolase A, an essential enzyme in glycolysis. We further show that mTORC1-dependent Aldolase A translation repression in D1 neurons is mediated through upregulation of miR-34a-5p expression. Alcohol-mediated mTORC1 repression of Aldolase A translation in D1 neurons inhibits glycolysis in the NAc. Finally, we report that overexpression of miR-34a-5p in D1 NAc neurons increases, whereas systemic administration of L-lactate, the final product of glycolysis, attenuates excessive alcohol intake. Our data suggest that alcohol promotes paradoxical actions of mTORC1 on translation and glycolysis which in turn drive excessive alcohol use.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures


Update of
-
Paradoxical mTORC1-Dependent microRNA-mediated Translation Repression in the Nucleus Accumbens of Mice Consuming Alcohol Attenuates Glycolysis.bioRxiv [Preprint]. 2025 Jan 28:2023.11.29.569312. doi: 10.1101/2023.11.29.569312. bioRxiv. 2025. Update in: Nat Commun. 2025 Jul 14;16(1):6116. doi: 10.1038/s41467-025-60337-9. PMID: 38076984 Free PMC article. Updated. Preprint.
Similar articles
-
Paradoxical mTORC1-Dependent microRNA-mediated Translation Repression in the Nucleus Accumbens of Mice Consuming Alcohol Attenuates Glycolysis.bioRxiv [Preprint]. 2025 Jan 28:2023.11.29.569312. doi: 10.1101/2023.11.29.569312. bioRxiv. 2025. Update in: Nat Commun. 2025 Jul 14;16(1):6116. doi: 10.1038/s41467-025-60337-9. PMID: 38076984 Free PMC article. Updated. Preprint.
-
The First Alcohol Drink Triggers mTORC1-Dependent Synaptic Plasticity in Nucleus Accumbens Dopamine D1 Receptor Neurons.J Neurosci. 2016 Jan 20;36(3):701-13. doi: 10.1523/JNEUROSCI.2254-15.2016. J Neurosci. 2016. PMID: 26791202 Free PMC article.
-
Brain-specific inhibition of mTORC1 eliminates side effects resulting from mTORC1 blockade in the periphery and reduces alcohol intake in mice.Nat Commun. 2021 Jul 27;12(1):4407. doi: 10.1038/s41467-021-24567-x. Nat Commun. 2021. PMID: 34315870 Free PMC article.
-
Non-invasive and invasive brain stimulation in alcohol use disorders: A critical review of selected human evidence and methodological considerations to guide future research.Compr Psychiatry. 2021 Aug;109:152257. doi: 10.1016/j.comppsych.2021.152257. Epub 2021 Jul 3. Compr Psychiatry. 2021. PMID: 34246194
-
Effectiveness of brief alcohol interventions in primary care populations.Cochrane Database Syst Rev. 2018 Feb 24;2(2):CD004148. doi: 10.1002/14651858.CD004148.pub4. Cochrane Database Syst Rev. 2018. PMID: 29476653 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

