This is a preprint.
Gene set optimization for cancer transcriptomics using sparse principal component analysis
- PMID: 40661635
- PMCID: PMC12258712
- DOI: 10.1101/2025.05.21.655279
Gene set optimization for cancer transcriptomics using sparse principal component analysis
Abstract
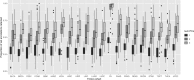
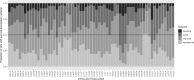
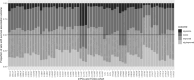
A common approach for exploring pathway dysregulation in cancer involves the gene set or pathway analysis of tumor transcriptomic data. Unfortunately, the effectiveness of cancer gene set testing is limited by the fact that most gene set collections model gene activity in normal tissue, which can differ significantly from gene activity found within tumors. To address this challenge, we have developed a bioinformatics approach based on sparse principal component analysis (PCA) for optimizing existing gene set collections to reflect the pattern of gene activity in dysplastic tissue and have used this technique to optimize the Molecular Signatures Database (MSigDB) Hallmark collection for 21 solid human cancers profiled via bulk RNA-seq by The Tumor Genome Atlas (TCGA). Demonstrating the biological utility of our approach, the average survival association of gene set members is improved after optimization for nearly all cancer types and Hallmark gene sets.
Conflict of interest statement
Conflict of interests The authors have no conflicts of interest to declare.
Figures
Similar articles
-
Can a Liquid Biopsy Detect Circulating Tumor DNA With Low-passage Whole-genome Sequencing in Patients With a Sarcoma? A Pilot Evaluation.Clin Orthop Relat Res. 2025 Jan 1;483(1):39-48. doi: 10.1097/CORR.0000000000003161. Epub 2024 Jun 21. Clin Orthop Relat Res. 2025. PMID: 38905450
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2021 Apr 19;4(4):CD011535. doi: 10.1002/14651858.CD011535.pub4. Cochrane Database Syst Rev. 2021. Update in: Cochrane Database Syst Rev. 2022 May 23;5:CD011535. doi: 10.1002/14651858.CD011535.pub5. PMID: 33871055 Free PMC article. Updated.
-
Impact of residual disease as a prognostic factor for survival in women with advanced epithelial ovarian cancer after primary surgery.Cochrane Database Syst Rev. 2022 Sep 26;9(9):CD015048. doi: 10.1002/14651858.CD015048.pub2. Cochrane Database Syst Rev. 2022. PMID: 36161421 Free PMC article.
-
Short-Term Memory Impairment.2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 31424720 Free Books & Documents.
-
Cost-effectiveness of using prognostic information to select women with breast cancer for adjuvant systemic therapy.Health Technol Assess. 2006 Sep;10(34):iii-iv, ix-xi, 1-204. doi: 10.3310/hta10340. Health Technol Assess. 2006. PMID: 16959170
References
-
- Ashburner Michael, Ball Catherine A., Blake Judith A., Botstein David, Butler Heather, Michael Cherry J., Davis Allan P., Dolinski Kara, Dwight Selina S., Eppig Janan T., Harris Midori A., Hill David P., Laurie Issel-Tarver Andrew Kasarskis, Lewis Suzanna, Matese John C., Richardson Joel E., Ringwald Martin, Rubin Gerald M., and Sherlock Gavin. Gene ontology: tool for the unification of biology. Nature Genetics, 25(1):25–29, May 2000. - PMC - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
