Single-cell RNA-sequencing highlights a curtailed NK cell function in convalescent COVID-19 pregnant women
- PMID: 40661959
- PMCID: PMC12257036
- DOI: 10.3389/fimmu.2025.1560391
Single-cell RNA-sequencing highlights a curtailed NK cell function in convalescent COVID-19 pregnant women
Abstract
Introduction: During gestation the immune system undergoes dramatic remodelling to protect the maternal-fetal dyad from infections whilst also preventing fetal rejection. We investigated how SARS-CoV-2 modifies the immune landscape during infection and in recovered pregnant women.
Methods: We immunophenotyped our two independent geographical cohorts using a 14-colour flow cytometry panel (surface and intracellular staining). We estimated cytokines and SARS-CoV-2 IgG antibodies in validation cohort using a multiplexd flow cytometry panel. Single-cell RNA sequencing (scRNA-seq) was performed using a Chromium Single Cell 3' Gel Bead Chip and Library Kit from 10x Genomics (Drop-seq method). Furthermore, we estimated the cytotoxic functions of natural killer (NK) cells by flow cytometry using surface and intracellular staining.
Results: Using two independent geographical cohorts, we identified that NK cells had a sustained reduction during active infection and after recovery. Further, scRNA-seq data revealed that infection with SARS-CoV-2 rewired the gene expression profile of NK, monocytes, CD4+, CD8+ effector T cells and antibody producing B cells in convalescent pregnant women. Several gene pathways associated with cytotoxic function, interferon signalling type I & II, and pro- and anti-inflammatory functions in NK and CD8+ cytotoxic T cells were attenuated in recovered pregnant patients compared with healthy pregnancies. We validated our scRNA-seq of NK cells from convalescent pregnant women and confirmed that NK cells had diminished levels of cytotoxic proteins; perforin, CD122 and granzyme B.
Discussion: Overall, our study uncovers that SARS-CoV-2 infection deranges the adaptive immune response in pregnant women even after recovery and may contribute to post-COVID19 sequalae of symptoms.
Keywords: COVID-19; NK cells; PBMCs; ScRNA-seq; cytotoxic cells; immunophenotyping and cytokines; pregnancy.
Copyright © 2025 Salker, Abd Aziz, Prodan, Yang, Lankapalli, Lazar, Shafiee, Kocak, Ganesan, Pal, Khalid, Kasim, Kalok, Fuad, Bissinger, Ganzenmueller, Iftner, Kagan, Ossowski, Casadei, Brucker, Riess and Singh.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures




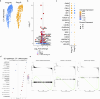



References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

