This is a preprint.
CDH1 loss remodels gene expression and lineage identity in human mammary epithelial cells
- PMID: 40667170
- PMCID: PMC12262285
- DOI: 10.1101/2025.06.20.660633
CDH1 loss remodels gene expression and lineage identity in human mammary epithelial cells
Abstract
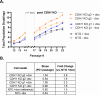
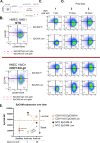
Invasive lobular carcinoma (ILC) is a common subtype of breast cancer that is defined in part by genetic loss of CDH1 caused by mutation or deletion, leading to loss of cell adhesion protein E-cadherin in >90% of ILC. Genetic loss of CDH1 is an early event in ILC oncogenesis, yet the mechanisms by which CDH1/E-cadherin acts as a tumor suppressor are not well understood. To study how early CDH1 loss drives ILC oncogenesis, we used a series of non-transformed human mammary epithelial cell (HMEC) models to target CDH1/E-cadherin, inhibiting extracellular E-cadherin signaling using antibodies versus modeling genetic CDH1 loss using siRNA or knockout via CRISPR/Cas9. Through transcriptome analyses across four HMEC models, we found that the mode of E-cadherin loss or suppression is critical for the subsequent phenotype. Antibody-mediated inhibition of cell-cell contacts induced gene signatures of epithelial-mesenchymal transition (EMT), consistent with the role of E-cadherin suppression during the EMT process. Conversely, genetic CDH1 loss - as in ILC oncogenesis - repressed EMT signatures, and instead remodeled gene expression toward a luminal epithelial phenotype. Using single cell transcriptomics and flow cytometry analyses of cell lineage markers, we found that genetic loss of CDH1 reprogrammed cells to a luminal progenitor-like phenotype. By isolating luminal versus basal cells prior to CDH1 knockout, we found that CDH1 loss led to remodeling of lineage identity in both populations, converging on a new lineage homeostasis with a luminal progenitor-like phenotype. Consistent with increased progenitor features, CDH1 loss enhanced proliferative capacity over the finite lifespan of the HMECs, highlighting a feature of early CDH1 loss that may contribute to clonal advantage during tumor initiation. Our findings support that inhibition of E-cadherin results in different transcriptional response compared to CDH1 loss, with the latter driving a transcriptional and phenotypic state characteristic of a luminal progenitor-like population, which offers new insight into early events in ILC oncogenesis.
Keywords: CDH1; E-cadherin; EMT; Invasive lobular carcinoma; breast cancer; etiology; mammary gland; oncogenesis.
Figures




References
-
- Grabenstetter A., Mohanty A.S., Rana S., Zehir A., Brannon A.R., D’Alfonso T.M., DeLair D.F., Tan L.K., Ross D.S., E-cadherin immunohistochemical expression in invasive lobular carcinoma of the breast: correlation with morphology and CDH1 somatic alterations, Hum Pathol 102 (2020) 44–53. 10.1016/j.humpath.2020.06.002. - DOI - PMC - PubMed
-
- Mathew A., Rajagopal P.S., Villgran V., Sandhu G.S., Jankowitz R.C., Jacob M., Rosenzweig M., Oesterreich S., Brufsky A., Distinct Pattern of Metastases in Patients with Invasive Lobular Carcinoma of the Breast, Geburtshilfe Frauenheilkd 77 (2017) 660–666. 10.1055/s-0043-109374. - DOI - PMC - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
